ING2 is upregulated in colon cancer and increases invasion by enhanced MMP13 expression
- PMID: 19437536
- PMCID: PMC2790157
- DOI: 10.1002/ijc.24437
ING2 is upregulated in colon cancer and increases invasion by enhanced MMP13 expression
Abstract
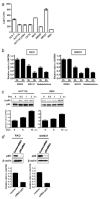
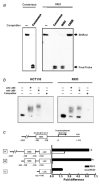
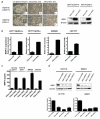
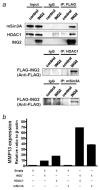
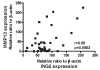
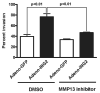
Inhibitor of growth 2 (ING2) is associated with chromatin remodeling and regulation of gene expression by binding to a methylated histone H3K4 residue and recruiting HDAC complexes to the region. The aim of our study is to investigate the regulation of ING2 expression and the clinical significance of upregulated ING2 in colon cancer. Here, we show that the ING2 mRNA level in colon cancer tissue increased to more than twice than that in normal mucosa in the 45% of colorectal cancer cases that we examined. A putative NF-kappaB binding site was found in the ING2 promoter region. We confirmed that NF-kappaB could bind to the ING2 promoter by EMSA and luciferase assays. Subsequent microarray analyses revealed that ING2 upregulates expression of matrix metalloproteinase 13 (MMP13), which enhances cancer invasion and metastasis. ING2 regulation of MMP13 expression was confirmed in both ING2 overexpression and knock down experiments. MMP13 expression was further induced by coexpression of ING2 with HDAC1 or with mSin3A, suggesting that the ING2-HDAC1-mSin3A complex members regulates expression of MMP13. In vitro invasion assay was performed to determine functional significance of ING2 upregulation. ING2 overexpressed cells exhibited greater invasive potential. Taken together, upregulation of ING2 was associated with colon cancer and MMP13-dependent cellular invasion, indicating that ING2 expression might be involved with cancer invasion and metastasis.
2009 UICC
Figures

Similar articles
-
ING2, a tumor associated gene, enhances PAI‑1 and HSPA1A expression with HDAC1 and mSin3A through the PHD domain and C‑terminal.Mol Med Rep. 2017 Nov;16(5):7367-7374. doi: 10.3892/mmr.2017.7553. Epub 2017 Sep 20. Mol Med Rep. 2017. PMID: 28944862 Free PMC article.
-
Decreased expression of ING2 gene and its clinicopathological significance in Chinese NSCLC patients.Neoplasma. 2014;61(4):468-75. doi: 10.4149/neo_2014_058. Neoplasma. 2014. PMID: 24712846
-
Knockdown of inhibitor of growth protein 2 inhibits cell invasion and enhances chemosensitivity to 5-FU in human gastric cancer cells.Dig Dis Sci. 2013 Nov;58(11):3189-97. doi: 10.1007/s10620-013-2796-5. Epub 2013 Jul 18. Dig Dis Sci. 2013. PMID: 23864195
-
ING1 and ING2: multifaceted tumor suppressor genes.Cell Mol Life Sci. 2013 Oct;70(20):3753-72. doi: 10.1007/s00018-013-1270-z. Epub 2013 Feb 15. Cell Mol Life Sci. 2013. PMID: 23412501 Free PMC article. Review.
-
ING proteins as potential anticancer drug targets.Curr Drug Targets. 2009 May;10(5):442-54. doi: 10.2174/138945009788185059. Curr Drug Targets. 2009. PMID: 19442116 Free PMC article. Review.
Cited by
-
Exploiting ING2 Epigenetic Modulation as a Therapeutic Opportunity for Non-Small Cell Lung Cancer.Cancers (Basel). 2019 Oct 21;11(10):1601. doi: 10.3390/cancers11101601. Cancers (Basel). 2019. PMID: 31640185 Free PMC article. Review.
-
Targeted disruption of Ing2 results in defective spermatogenesis and development of soft-tissue sarcomas.PLoS One. 2010 Nov 19;5(11):e15541. doi: 10.1371/journal.pone.0015541. PLoS One. 2010. PMID: 21124965 Free PMC article.
-
Immunohistochemical analysis of MMP-13 and EMMPRIN in epithelial odontogenic lesions.Eur Arch Otorhinolaryngol. 2019 Nov;276(11):3203-3211. doi: 10.1007/s00405-019-05589-0. Epub 2019 Aug 22. Eur Arch Otorhinolaryngol. 2019. PMID: 31440816
-
Inhibitor of growth tumor suppressors in cancer progression.Cell Mol Life Sci. 2010 Jun;67(12):1987-99. doi: 10.1007/s00018-010-0312-z. Epub 2010 Mar 2. Cell Mol Life Sci. 2010. PMID: 20195696 Free PMC article. Review.
-
Multidimensional analysis to elucidate the possible mechanism of bone metastasis in breast cancer.BMC Cancer. 2023 Dec 8;23(1):1213. doi: 10.1186/s12885-023-11588-6. BMC Cancer. 2023. PMID: 38066539 Free PMC article.
References
-
- Jemal A, Siegel R, Ward E, Hao Y, Xu J, Murray T, Thun MJ. Cancer statistics, 2008. CA Cancer. J Clin. 2008;58:71–96. - PubMed
-
- Shimada Y, Saito A, Suzuki M, Takahashi E, Horie M. Cloning of a novel gene (ING1L) homologous to ING1, a candidate tumor suppressor. Cytogenet Cell Genet. 1998;83:232–5. - PubMed
-
- Garkavtsev I, Kazarov A, Gudkov A, Riabowol K. Suppression of the novel growth inhibitor p33ING1 promotes neoplastic transformation. Nat Genet. 1996;14:415–20. - PubMed
-
- Garkavtsev I, Grigorian IA, Ossovskaya VS, Chernov MV, Chumakov PM, Gudkov AV. The candidate tumour suppressor p33ING1 cooperates with p53 in cell growth control. Nature. 1998;391:295–8. - PubMed
-
- Feng X, Hara Y, Riabowol K. Different HATS of the ING1 gene family. Trends Cell Biol. 2002;12:532–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

