Structure and site-specific recognition of histone H3 by the PHD finger of human autoimmune regulator
- PMID: 19446523
- PMCID: PMC2923636
- DOI: 10.1016/j.str.2009.02.017
Structure and site-specific recognition of histone H3 by the PHD finger of human autoimmune regulator
Abstract
Human autoimmune regulator (AIRE) functions to control thymic expression of tissue-specific antigens via sequence-specific histone H3 recognition by its plant homeodomain (PHD) finger. Mutations in the AIRE PHD finger have been linked to autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy (APECED). Here we report the three-dimensional solution structure of the first PHD finger of human AIRE bound to a histone H3 peptide. The structure reveals a detailed network of interactions between the protein and the amino-terminal residues of histone H3, and particularly key electrostatic interactions of a conserved aspartic acid 297 in AIRE with the unmodified lysine 4 of histone H3 (H3K4). NMR binding study with H3 peptides carrying known posttranslational modifications flanking H3K4 confirms that transcriptional regulation by AIRE through its interactions with histone H3 is confined to the first N-terminal eight residues in H3. Our study offers a molecular explanation for the APECED mutations and helps define a subclass of the PHD finger family proteins that recognize histone H3 in a sequence-specific manner.
Figures

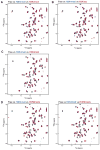


References
-
- Aasland R, Gibson TJ, Stewart AF. The PHD finger: implications for chromatin-mediated transcriptional regulation. Trends Biochem Sci. 1995;20:56–59. - PubMed
-
- Anderson MS, Venanzi ES, Klein L, Chen Z, Berzins SP, Turley SJ, von Boehmer H, Bronson R, Dierich A, Benoist C, Mathis D. Projection of an immunological self shadow within the thymus by the aire protein. Science. 2002;298:1395–1401. - PubMed
-
- Aravind L, Iyer L. Provenance of SET-domain histone methyl-transferases through duplication of a simple structural unit. Cell Cycle. 2003;2:369–376. - PubMed
-
- Aravind L, Iyer LM, Koonin EV. Comparative genomics and structural biology of the molecular innovations of eukaryotes. Curr Opin Struct Biol. 2006;16:409–419. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

