Apical polarity protein PrkCi is necessary for maintenance of spinal cord precursors in zebrafish
- PMID: 19449304
- PMCID: PMC2829461
- DOI: 10.1002/dvdy.21970
Apical polarity protein PrkCi is necessary for maintenance of spinal cord precursors in zebrafish
Abstract
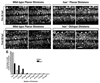
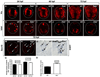
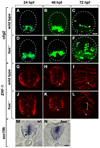
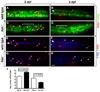
During development, neural precursors divide to produce new precursors and cells that differentiate as neurons and glia. In Drosophila, apicobasal polarity and orientation of the mitotic spindle play important roles in specifying the progeny of neural precursors for different fates. We examined orientation of zebrafish spinal cord precursors using time-lapse imaging and tested the function of protein kinase C, iota (PrkCi), a member of the Par complex of proteins necessary for apicobasal polarity in the nervous system. We found that nearly all precursors divide within the plane of the neuroepithelium of wild-type embryos even when they must produce cells that have different fates. In the absence of PrkCi function, neural precursor divisions become oblique during late embryogenesis and excess oligodendrocytes form concomitant with loss of dividing cells. We conclude that PrkCi function and planar divisions are necessary for asymmetric, self-renewing division of spinal cord precursors.
(c) 2009 Wiley-Liss, Inc.
Figures


Similar articles
-
Positional cloning of heart and soul reveals multiple roles for PKC lambda in zebrafish organogenesis.Curr Biol. 2001 Oct 2;11(19):1492-502. doi: 10.1016/s0960-9822(01)00458-4. Curr Biol. 2001. PMID: 11591316
-
Notch-regulated oligodendrocyte specification from radial glia in the spinal cord of zebrafish embryos.Dev Dyn. 2008 Aug;237(8):2081-9. doi: 10.1002/dvdy.21620. Dev Dyn. 2008. PMID: 18627107 Free PMC article.
-
Control of planar divisions by the G-protein regulator LGN maintains progenitors in the chick neuroepithelium.Nat Neurosci. 2007 Nov;10(11):1440-8. doi: 10.1038/nn1984. Epub 2007 Oct 14. Nat Neurosci. 2007. PMID: 17934458
-
Generating asymmetry: with and without self-renewal.Prog Mol Subcell Biol. 2007;45:143-78. doi: 10.1007/978-3-540-69161-7_7. Prog Mol Subcell Biol. 2007. PMID: 17585500 Review.
-
From cells to circuits: development of the zebrafish spinal cord.Prog Neurobiol. 2003 Apr;69(6):419-49. doi: 10.1016/s0301-0082(03)00052-2. Prog Neurobiol. 2003. PMID: 12880634 Review.
Cited by
-
The neuroepithelial basement membrane serves as a boundary and a substrate for neuron migration in the zebrafish hindbrain.Neural Dev. 2010 Mar 29;5:9. doi: 10.1186/1749-8104-5-9. Neural Dev. 2010. PMID: 20350296 Free PMC article.
-
Identification of Arhgef12 and Prkci as genetic modifiers of retinal dysplasia in the Crb1rd8 mouse model.PLoS Genet. 2022 Jun 8;18(6):e1009798. doi: 10.1371/journal.pgen.1009798. eCollection 2022 Jun. PLoS Genet. 2022. PMID: 35675330 Free PMC article.
-
Subnuclear development of the zebrafish habenular nuclei requires ER translocon function.Dev Biol. 2011 Dec 1;360(1):44-57. doi: 10.1016/j.ydbio.2011.09.003. Epub 2011 Sep 16. Dev Biol. 2011. PMID: 21945073 Free PMC article.
-
miR-219 regulates neural precursor differentiation by direct inhibition of apical par polarity proteins.Dev Cell. 2013 Nov 25;27(4):387-98. doi: 10.1016/j.devcel.2013.10.015. Epub 2013 Nov 14. Dev Cell. 2013. PMID: 24239515 Free PMC article.
-
Transgenic line with gal4 insertion useful to study morphogenesis of craniofacial perichondrium, vascular endothelium-associated cells, floor plate, and dorsal midline radial glia during zebrafish development.Dev Growth Differ. 2012 Feb;54(2):202-15. doi: 10.1111/j.1440-169X.2011.01322.x. Epub 2012 Feb 20. Dev Growth Differ. 2012. PMID: 22348745 Free PMC article.
References
-
- Aaku-Saraste E, Hellwig A, Huttner WB. Loss of occludin and functional tight junctions, but not ZO-1, during neural tube closure—remodeling of the neuroepithelium prior to neurogenesis. Dev Biol. 1996;180:664–679. - PubMed
-
- Afonso C, Henrique D. PAR3 acts as a molecular organizer to define the apical domain of chick neuroepithelial cells. J Cell Sci. 2006;119:4293–4304. - PubMed
-
- Betschinger J, Mechtler K, Knoblich JA. The Par complex directs asymmetric cell division by phosphorylating the cytoskeletal protein Lgl. Nature. 2003;422:326–330. - PubMed
-
- Buchman JJ, Tsai LH. Putting a notch in our understanding of nuclear migration. Cell. 2008;134:912–914. - PubMed
-
- Chenn A, McConnell SK. Cleavage orientation and the asymmetric inheritance of Notch1 immunoreactivity in mammalian neurogenesis. Cell. 1995;82:631–641. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

