The H3K27me3 demethylase JMJD3 contributes to the activation of the INK4A-ARF locus in response to oncogene- and stress-induced senescence
- PMID: 19451217
- PMCID: PMC2685535
- DOI: 10.1101/gad.510809
The H3K27me3 demethylase JMJD3 contributes to the activation of the INK4A-ARF locus in response to oncogene- and stress-induced senescence
Abstract
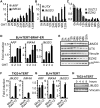
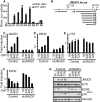
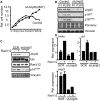
The tumor suppressor proteins p16INK4A and p14ARF, encoded by the INK4A-ARF locus, are key regulators of cellular senescence. The locus is epigenetically silenced by the repressive H3K27me3 mark in normally growing cells, but becomes activated in response to oncogenic stress. Here, we show that expression of the histone H3 Lys 27 (H3K27) demethylase JMJD3 is induced upon activation of the RAS-RAF signaling pathway. JMJD3 is recruited to the INK4A-ARF locus and contributes to the transcriptional activation of p16INK4A in human diploid fibroblasts. Additionally, inhibition of Jmjd3 expression in mouse embryonic fibroblasts results in suppression of p16Ink4a and p19Arf expression and in their immortalization.
Figures


References
-
- Agger K, Cloos PA, Christensen J, Pasini D, Rose S, Rappsilber J, Issaeva I, Canaani E, Salcini AE, Helin K. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature. 2007;449:731–734. - PubMed
-
- Ameyar-Zazoua M, Wisniewska MB, Bakiri L, Wagner EF, Yaniv M, Weitzman JB. AP-1 dimers regulate transcription of the p14/p19ARF tumor suppressor gene. Oncogene. 2005;24:2298–2306. - PubMed
-
- Basso K, Margolin AA, Stolovitzky G, Klein U, Dalla-Favera R, Califano A. Reverse engineering of regulatory networks in human B cells. Nat Genet. 2005;37:382–390. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
