Universal partitioning of the hierarchical fold network of 50-residue segments in proteins
- PMID: 19454039
- PMCID: PMC2693521
- DOI: 10.1186/1472-6807-9-34
Universal partitioning of the hierarchical fold network of 50-residue segments in proteins
Abstract
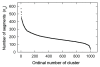
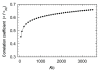
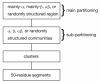
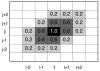
Background: Several studies have demonstrated that protein fold space is structured hierarchically and that power-law statistics are satisfied in relation between the numbers of protein families and protein folds (or superfamilies). We examined the internal structure and statistics in the fold space of 50 amino-acid residue segments taken from various protein folds. We used inter-residue contact patterns to measure the tertiary structural similarity among segments. Using this similarity measure, the segments were classified into a number (Kc) of clusters. We examined various Kc values for the clustering. The special resolution to differentiate the segment tertiary structures increases with increasing Kc. Furthermore, we constructed networks by linking structurally similar clusters.
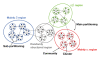
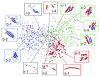
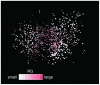
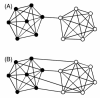
Results: The network was partitioned persistently into four regions for Kc >or= 1000. This main partitioning is consistent with results of earlier studies, where similar partitioning was reported in classifying protein domain structures. Furthermore, the network was partitioned naturally into several dozens of sub-networks (i.e., communities). Therefore, intra-sub-network clusters were mutually connected with numerous links, although inter-sub-network ones were rarely done with few links. For Kc >or= 1000, the major sub-networks were about 40; the contents of the major sub-networks were conserved. This sub-partitioning is a novel finding, suggesting that the network is structured hierarchically: Segments construct a cluster, clusters form a sub-network, and sub-networks constitute a region. Additionally, the network was characterized by non-power-law statistics, which is also a novel finding.
Conclusion: Main findings are: (1) The universe of 50 residue segments found here was characterized by non-power-law statistics. Therefore, the universe differs from those ever reported for the protein domains. (2) The 50-residue segments were partitioned persistently and universally into some dozens (ca. 40) of major sub-networks, irrespective of the number of clusters. (3) These major sub-networks encompassed 90% of all segments. Consequently, the protein tertiary structure is constructed using the dozens of elements (sub-networks).
Figures

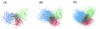
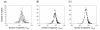


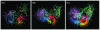
Similar articles
-
Atomic interaction networks in the core of protein domains and their native folds.PLoS One. 2010 Feb 23;5(2):e9391. doi: 10.1371/journal.pone.0009391. PLoS One. 2010. PMID: 20186337 Free PMC article.
-
Cross-over between discrete and continuous protein structure space: insights into automatic classification and networks of protein structures.PLoS Comput Biol. 2009 Mar;5(3):e1000331. doi: 10.1371/journal.pcbi.1000331. Epub 2009 Mar 27. PLoS Comput Biol. 2009. PMID: 19325884 Free PMC article.
-
Exploring dynamics of protein structure determination and homology-based prediction to estimate the number of superfamilies and folds.BMC Struct Biol. 2006 Mar 20;6:6. doi: 10.1186/1472-6807-6-6. BMC Struct Biol. 2006. PMID: 16549009 Free PMC article.
-
Intra and inter-molecular communications through protein structure network.Curr Protein Pept Sci. 2009 Apr;10(2):146-60. doi: 10.2174/138920309787847590. Curr Protein Pept Sci. 2009. PMID: 19355982 Review.
-
The structure of the protein universe and genome evolution.Nature. 2002 Nov 14;420(6912):218-23. doi: 10.1038/nature01256. Nature. 2002. PMID: 12432406 Review.
References
-
- Murzin AG, Brenner SE, Hubbard T, Chothia C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J Mol Biol. 1995;247:536–540. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

