Operator-assisted harvesting of protein crystals using a universal micromanipulation robot
- PMID: 19461845
- PMCID: PMC2483483
- DOI: 10.1107/S0021889807012149
Operator-assisted harvesting of protein crystals using a universal micromanipulation robot
Abstract
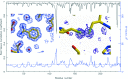
High-throughput crystallography has reached a level of automation where complete computer-assisted robotic crystallization pipelines are capable of cocktail preparation, crystallization plate setup, and inspection and interpretation of results. While mounting of crystal pins, data collection and structure solution are highly automated, crystal harvesting and cryocooling remain formidable challenges towards full automation. To address the final frontier in achieving fully automated high-throughput crystallography, the prototype of an anthropomorphic six-axis universal micromanipulation robot (UMR) has been designed and tested; this UMR is capable of operator-assisted harvesting and cryoquenching of protein crystals as small as 10 microm from a variety of 96-well plates. The UMR is equipped with a versatile tool exchanger providing full operational flexibility. Trypsin crystals harvested and cryoquenched using the UMR have yielded a 1.5 A structure demonstrating the feasibility of robotic protein crystal harvesting.
Figures




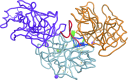

Similar articles
-
Automated robotic harvesting of protein crystals-addressing a critical bottleneck or instrumentation overkill?J Struct Funct Genomics. 2007 Dec;8(4):145-52. doi: 10.1007/s10969-007-9031-6. Epub 2007 Oct 27. J Struct Funct Genomics. 2007. PMID: 17965947 Review.
-
First experiences with semi-autonomous robotic harvesting of protein crystals.J Struct Funct Genomics. 2011 Jul;12(2):77-82. doi: 10.1007/s10969-011-9103-5. Epub 2011 Mar 23. J Struct Funct Genomics. 2011. PMID: 21431335
-
REACH: Robotic Equipment for Automated Crystal Harvesting using a six-axis robot arm and a micro-gripper.Acta Crystallogr D Biol Crystallogr. 2013 Mar;69(Pt 3):381-7. doi: 10.1107/S0907444912048019. Epub 2013 Feb 16. Acta Crystallogr D Biol Crystallogr. 2013. PMID: 23519413
-
CrystalDirect™: a novel approach for automated crystal harvesting based on photoablation of thin films.Methods Mol Biol. 2014;1091:197-203. doi: 10.1007/978-1-62703-691-7_14. Methods Mol Biol. 2014. PMID: 24203334
-
Life in the fast lane for protein crystallization and X-ray crystallography.Prog Biophys Mol Biol. 2005 Jul;88(3):359-86. doi: 10.1016/j.pbiomolbio.2004.07.011. Prog Biophys Mol Biol. 2005. PMID: 15652250 Review.
Cited by
-
Acoustic transfer of protein crystals from agarose pedestals to micromeshes for high-throughput screening.Acta Crystallogr D Biol Crystallogr. 2015 Jan 1;71(Pt 1):94-103. doi: 10.1107/S1399004714013728. Epub 2015 Jan 1. Acta Crystallogr D Biol Crystallogr. 2015. PMID: 25615864 Free PMC article.
-
Lessons from high-throughput protein crystallization screening: 10 years of practical experience.Expert Opin Drug Discov. 2011 May;6(5):465-80. doi: 10.1517/17460441.2011.566857. Epub 2011 Mar 22. Expert Opin Drug Discov. 2011. PMID: 22646073 Free PMC article.
-
The solvent component of macromolecular crystals.Acta Crystallogr D Biol Crystallogr. 2015 May;71(Pt 5):1023-38. doi: 10.1107/S1399004715006045. Epub 2015 Apr 30. Acta Crystallogr D Biol Crystallogr. 2015. PMID: 25945568 Free PMC article. Review.
-
Development of an alternative approach to protein crystallization.J Struct Funct Genomics. 2007 Dec;8(4):193-8. doi: 10.1007/s10969-007-9034-3. Epub 2007 Nov 24. J Struct Funct Genomics. 2007. PMID: 18038192
-
Using sound pulses to solve the crystal-harvesting bottleneck.Acta Crystallogr D Struct Biol. 2018 Oct 1;74(Pt 10):986-999. doi: 10.1107/S2059798318011506. Epub 2018 Oct 2. Acta Crystallogr D Struct Biol. 2018. PMID: 30289409 Free PMC article.
References
-
- Abagyan, R., Totrov, M. & Kuznetsov, D. (1994). J. Comput. Chem. 15, 488–506.
-
- Balschmidt, P., Hansen, F. B., Dodson, E. J., Dodson, G. G. & Korber, F. (1991). Acta Cryst. B47, 975–986. - PubMed
-
- Beteva, A. et al. (2006). Acta Cryst. D62, 1162–1169. - PubMed
-
- Blundell, T. L., Jhoti, H. & Abell, C. (2002). Nat. Rev. Drug Discov. 1, 45–54. - PubMed
-
- Bruker (2004). PROTEUM Bruker AXS Inc., Madison, Wisconsin, USA.
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
