Deaminase-independent inhibition of parvoviruses by the APOBEC3A cytidine deaminase
- PMID: 19461882
- PMCID: PMC2678267
- DOI: 10.1371/journal.ppat.1000439
Deaminase-independent inhibition of parvoviruses by the APOBEC3A cytidine deaminase
Abstract
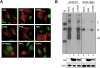
The APOBEC3 proteins form a multigene family of cytidine deaminases with inhibitory activity against viruses and retrotransposons. In contrast to APOBEC3G (A3G), APOBEC3A (A3A) has no effect on lentiviruses but dramatically inhibits replication of the parvovirus adeno-associated virus (AAV). To study the contribution of deaminase activity to the antiviral activity of A3A, we performed a comprehensive mutational analysis of A3A. By mutation of non-conserved residues, we found that regions outside of the catalytic active site contribute to both deaminase and antiviral activities. Using A3A point mutants and A3A/A3G chimeras, we show that deaminase activity is not required for inhibition of recombinant AAV production. We also found that deaminase-deficient A3A mutants block replication of both wild-type AAV and the autonomous parvovirus minute virus of mice (MVM). In addition, we identify specific residues of A3A that confer activity against AAV when substituted into A3G. In summary, our results demonstrate that deaminase activity is not necessary for the antiviral activity of A3A against parvoviruses.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

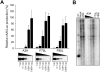




Similar articles
-
Structure-function analyses point to a polynucleotide-accommodating groove essential for APOBEC3A restriction activities.J Virol. 2011 Feb;85(4):1765-76. doi: 10.1128/JVI.01651-10. Epub 2010 Dec 1. J Virol. 2011. PMID: 21123384 Free PMC article.
-
Innate immune signaling induces high levels of TC-specific deaminase activity in primary monocyte-derived cells through expression of APOBEC3A isoforms.J Biol Chem. 2010 Sep 3;285(36):27753-66. doi: 10.1074/jbc.M110.102822. Epub 2010 Jul 8. J Biol Chem. 2010. PMID: 20615867 Free PMC article.
-
Functional domain organization of human APOBEC3G.Virology. 2008 Sep 15;379(1):118-24. doi: 10.1016/j.virol.2008.06.013. Epub 2008 Jul 18. Virology. 2008. PMID: 18639915 Free PMC article.
-
Role of the single deaminase domain APOBEC3A in virus restriction, retrotransposition, DNA damage and cancer.J Gen Virol. 2016 Jan;97(1):1-17. doi: 10.1099/jgv.0.000320. Epub 2015 Oct 20. J Gen Virol. 2016. PMID: 26489798 Free PMC article. Review.
-
Cytidine deamination and resistance to retroviral infection: towards a structural understanding of the APOBEC proteins.Virology. 2005 Apr 10;334(2):147-53. doi: 10.1016/j.virol.2005.01.038. Virology. 2005. PMID: 15780864 Review.
Cited by
-
APOBEC3 Interference during Replication of Viral Genomes.Viruses. 2015 Jun 11;7(6):2999-3018. doi: 10.3390/v7062757. Viruses. 2015. PMID: 26110583 Free PMC article. Review.
-
Infection of Bronchial Epithelial Cells by the Human Adenoviruses A12, B3, and C2 Differently Regulates the Innate Antiviral Effector APOBEC3B.J Virol. 2021 Jun 10;95(13):e0241320. doi: 10.1128/JVI.02413-20. Epub 2021 Jun 10. J Virol. 2021. PMID: 33853956 Free PMC article.
-
Lentivirus restriction by diverse primate APOBEC3A proteins.Virology. 2013 Jul 20;442(1):82-96. doi: 10.1016/j.virol.2013.04.002. Epub 2013 May 4. Virology. 2013. PMID: 23648232 Free PMC article.
-
Insights into the Structures and Multimeric Status of APOBEC Proteins Involved in Viral Restriction and Other Cellular Functions.Viruses. 2021 Mar 17;13(3):497. doi: 10.3390/v13030497. Viruses. 2021. PMID: 33802945 Free PMC article. Review.
-
Human Papillomavirus 16 E7 Stabilizes APOBEC3A Protein by Inhibiting Cullin 2-Dependent Protein Degradation.J Virol. 2018 Mar 14;92(7):e01318-17. doi: 10.1128/JVI.01318-17. Print 2018 Apr 1. J Virol. 2018. PMID: 29367246 Free PMC article.
References
-
- Jarmuz A, Chester A, Bayliss J, Gisbourne J, Dunham I, et al. An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics. 2002;79:285–296. - PubMed
-
- Zhang J, Webb DM. Rapid evolution of primate antiviral enzyme APOBEC3G. Hum Mol Genet. 2004;13:1785–1791. - PubMed
-
- Wedekind JE, Dance GS, Sowden MP, Smith HC. Messenger RNA editing in mammals: new members of the APOBEC family seeking roles in the family business. Trends Genet. 2003;19:207–216. - PubMed
-
- Harris RS, Liddament MT. Retroviral restriction by APOBEC proteins. Nat Rev Immunol. 2004;4:868–877. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

