RNA interference targeting leucine aminopeptidase blocks hatching of Schistosoma mansoni eggs
- PMID: 19463860
- PMCID: PMC2705689
- DOI: 10.1016/j.molbiopara.2009.05.002
RNA interference targeting leucine aminopeptidase blocks hatching of Schistosoma mansoni eggs
Abstract
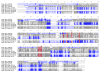
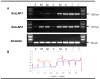
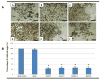
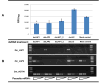
Schistosoma mansoni leucine aminopeptidase (LAP) is thought to play a central role in hatching of the miracidium from the schistosome egg. We identified two discrete LAPs genes in the S. mansoni genome, and their orthologs in S. japonicum. The similarities in sequence and exon/intron structure of the two genes, LAP1 and LAP2, suggest that they arose by gene duplication and that this occurred before separation of the mansoni and japonicum lineages. The SmLAP1 and SmLAP2 genes have different expression patterns in diverse stages of the cycle; whereas both are equally expressed in the blood dwelling stages (schistosomules and adult), SmLAP2 expression was higher in free living larval (miracidia) and in parasitic intra-snail (sporocysts) stages. We investigated the role of each enzyme in hatching of schistosome eggs and the early stages of schistosome development by RNA interference (RNAi). Using RNAi, we observed marked and specific reduction of mRNAs, along with a loss of exopeptidase activity in soluble parasite extracts against the diagnostic substrate l-leucine-7-amido-4-methylcoumarin hydroxide. Strikingly, knockdown of either SmLAP1 or SmLAP2, or both together, was accompanied by >or=80% inhibition of hatching of schistosome eggs showing that both enzymes are important to the escape of miracidia from the egg. The methods employed here refine the utility of RNAi for functional genomics studies in helminth parasites and confirm these can be used to identify potential drug targets, in this case schistosome aminopeptidases.
Figures


Similar articles
-
Expression, purification and characterization of two leucine aminopeptidases of the blood fluke, Schistosoma mansoni.Mol Biochem Parasitol. 2018 Jan;219:17-23. doi: 10.1016/j.molbiopara.2017.11.006. Epub 2017 Nov 21. Mol Biochem Parasitol. 2018. PMID: 29169803
-
Leucine aminopeptidase and hatching of Schistosoma mansoni eggs.J Parasitol. 1986 Aug;72(4):507-11. J Parasitol. 1986. PMID: 3783344
-
Leucine aminopeptidase of the human blood flukes, Schistosoma mansoni and Schistosoma japonicum.Int J Parasitol. 2004 May;34(6):703-14. doi: 10.1016/j.ijpara.2004.01.008. Int J Parasitol. 2004. PMID: 15111092
-
Progress with schistosome transgenesis.Mem Inst Oswaldo Cruz. 2011 Nov;106(7):785-93. doi: 10.1590/s0074-02762011000700002. Mem Inst Oswaldo Cruz. 2011. PMID: 22124549 Free PMC article. Review.
-
Culture for genetic manipulation of developmental stages of Schistosoma mansoni.Parasitology. 2010 Mar;137(3):451-62. doi: 10.1017/S0031182009991211. Epub 2009 Sep 21. Parasitology. 2010. PMID: 19765348 Free PMC article. Review.
Cited by
-
Transfection of Platyhelminthes.Biomed Res Int. 2015;2015:206161. doi: 10.1155/2015/206161. Epub 2015 May 18. Biomed Res Int. 2015. PMID: 26090388 Free PMC article. Review.
-
The tumorigenic liver fluke Opisthorchis viverrini--multiple pathways to cancer.Trends Parasitol. 2012 Oct;28(10):395-407. doi: 10.1016/j.pt.2012.07.006. Epub 2012 Sep 1. Trends Parasitol. 2012. PMID: 22947297 Free PMC article. Review.
-
RNA interference in Schistosoma mansoni schistosomula: selectivity, sensitivity and operation for larger-scale screening.PLoS Negl Trop Dis. 2010 Oct 19;4(10):e850. doi: 10.1371/journal.pntd.0000850. PLoS Negl Trop Dis. 2010. PMID: 20976050 Free PMC article.
-
Vesicle-based secretion in schistosomes: Analysis of protein and microRNA (miRNA) content of exosome-like vesicles derived from Schistosoma mansoni.Sci Rep. 2018 Feb 19;8(1):3286. doi: 10.1038/s41598-018-21587-4. Sci Rep. 2018. PMID: 29459722 Free PMC article.
-
A portrait of the transcriptome of the neglected trematode, Fasciola gigantica--biological and biotechnological implications.PLoS Negl Trop Dis. 2011 Feb 1;5(2):e1004. doi: 10.1371/journal.pntd.0001004. PLoS Negl Trop Dis. 2011. PMID: 21408104 Free PMC article.
References
-
- King CH, Dickman K, Tisch DJ. Reassessment of the cost of chronic helmintic infection: a meta-analysis of disability-related outcomes in endemic schistosomiasis. Lancet. 2005;365(9470):1561–9. - PubMed
-
- Gryseels B, et al. Human schistosomiasis. Lancet. 2006;368(9541):1106–18. - PubMed
-
- Pearce EJ, MacDonald AS. The immunobiology of schistosomiasis. Nat Rev Immunol. 2002;2(7):499–511. - PubMed
-
- Kloetzel K. A collagenaselike substance produced by eggs of Schistosoma mansoni. J Parasitol. 1968;54(1):177–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

