Characterization of a gene cluster involved in 4-chlorocatechol degradation by Pseudomonas reinekei MT1
- PMID: 19465655
- PMCID: PMC2715737
- DOI: 10.1128/JB.00331-09
Characterization of a gene cluster involved in 4-chlorocatechol degradation by Pseudomonas reinekei MT1
Abstract
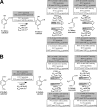
Pseudomonas reinekei MT1 has previously been reported to degrade 4- and 5-chlorosalicylate by a pathway with 4-chlorocatechol, 3-chloromuconate, 4-chloromuconolactone, and maleylacetate as intermediates, and a gene cluster channeling various salicylates into an intradiol cleavage route has been reported. We now report that during growth on 5-chlorosalicylate, besides a novel (chloro)catechol 1,2-dioxygenase, C12O(ccaA), a novel (chloro)muconate cycloisomerase, MCI(ccaB), which showed features not yet reported, was induced. This cycloisomerase, which was practically inactive with muconate, evolved for the turnover of 3-substituted muconates and transforms 3-chloromuconate into equal amounts of cis-dienelactone and protoanemonin, suggesting that it is a functional intermediate between chloromuconate cycloisomerases and muconate cycloisomerases. The corresponding genes, ccaA (C12O(ccaA)) and ccaB (MCI(ccaB)), were located in a 5.1-kb genomic region clustered with genes encoding trans-dienelactone hydrolase (ccaC) and maleylacetate reductase (ccaD) and a putative regulatory gene, ccaR, homologous to regulators of the IclR-type family. Thus, this region includes genes sufficient to enable MT1 to transform 4-chlorocatechol to 3-oxoadipate. Phylogenetic analysis showed that C12O(ccaA) and MCI(ccaB) are only distantly related to previously described catechol 1,2-dioxygenases and muconate cycloisomerases. Kinetic analysis indicated that MCI(ccaB) and the previously identified C12O(salD), rather than C12O(ccaA), are crucial for 5-chlorosalicylate degradation. Thus, MT1 uses enzymes encoded by a completely novel gene cluster for degradation of chlorosalicylates, which, together with a gene cluster encoding enzymes for channeling salicylates into the ortho-cleavage pathway, form an effective pathway for 4- and 5-chlorosalicylate mineralization.
Figures
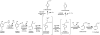
Similar articles
-
New bacterial pathway for 4- and 5-chlorosalicylate degradation via 4-chlorocatechol and maleylacetate in Pseudomonas sp. strain MT1.J Bacteriol. 2003 Dec;185(23):6790-800. doi: 10.1128/JB.185.23.6790-6800.2003. J Bacteriol. 2003. PMID: 14617643 Free PMC article.
-
A gene cluster involved in degradation of substituted salicylates via ortho cleavage in Pseudomonas sp. strain MT1 encodes enzymes specifically adapted for transformation of 4-methylcatechol and 3-methylmuconate.J Bacteriol. 2007 Mar;189(5):1664-74. doi: 10.1128/JB.01192-06. Epub 2006 Dec 15. J Bacteriol. 2007. PMID: 17172348 Free PMC article.
-
A new modified ortho cleavage pathway of 3-chlorocatechol degradation by Rhodococcus opacus 1CP: genetic and biochemical evidence.J Bacteriol. 2002 Oct;184(19):5282-92. doi: 10.1128/JB.184.19.5282-5292.2002. J Bacteriol. 2002. PMID: 12218013 Free PMC article.
-
Evolution of chlorocatechol catabolic pathways. Conclusions to be drawn from comparisons of lactone hydrolases.Biodegradation. 1994 Dec;5(3-4):301-21. doi: 10.1007/BF00696467. Biodegradation. 1994. PMID: 7765840 Review.
-
Transcriptional activation of the catechol and chlorocatechol operons: variations on a theme.Gene. 1998 Nov 26;223(1-2):257-67. doi: 10.1016/s0378-1119(98)00366-7. Gene. 1998. PMID: 9858745 Review.
Cited by
-
Novel degradation pathway of 4-chloro-2-aminophenol via 4-chlorocatechol in Burkholderia sp. RKJ 800.Environ Sci Pollut Res Int. 2014 Feb;21(3):2298-2304. doi: 10.1007/s11356-013-2167-y. Epub 2013 Sep 21. Environ Sci Pollut Res Int. 2014. PMID: 24057966
-
Managing gene expression in Pseudomonas simiae EGD-AQ6 for chloroaromatic compound degradation.Arch Microbiol. 2022 Jan 9;204(2):132. doi: 10.1007/s00203-021-02737-1. Arch Microbiol. 2022. PMID: 34999969
-
Biodegradation of chlorpropham and its major products by Bacillus licheniformis NKC-1.World J Microbiol Biotechnol. 2018 Jul 6;34(8):112. doi: 10.1007/s11274-018-2494-8. World J Microbiol Biotechnol. 2018. PMID: 29980862
-
The homogentisate and homoprotocatechuate central pathways are involved in 3- and 4-hydroxyphenylacetate degradation by Burkholderia xenovorans LB400.PLoS One. 2011 Mar 10;6(3):e17583. doi: 10.1371/journal.pone.0017583. PLoS One. 2011. PMID: 21423751 Free PMC article.
-
Diversity and Evolutionary Analysis of Iron-Containing (Type-III) Alcohol Dehydrogenases in Eukaryotes.PLoS One. 2016 Nov 28;11(11):e0166851. doi: 10.1371/journal.pone.0166851. eCollection 2016. PLoS One. 2016. PMID: 27893862 Free PMC article.
References
-
- Bertani, I., M. Kojic, and V. Venturi. 2001. Regulation of the p-hydroxybenzoic acid hydroxylase gene (pobA) in plant-growth-promoting Pseudomonas putida WCS358. Microbiology 1471611-1620. - PubMed
-
- Bhat, M. A., T. Ishida, K. Horiike, C. S. Vaidyanathan, and M. Nozaki. 1993. Purification of 3,5-dichlorocatechol 1,2-dioxygenase, a nonheme iron dioxygenase and a key enzyme in the biodegradation of a herbicide, 2,4-dichlorophenoxyacetic acid (2,4-D), from Pseudomonas cepacia CSV90. Arch. Biochem. Biophys. 300738-746. - PubMed
-
- Blasco, R., R.-M. Wittich, M. Mallavarapu, K. N. Timmis, and D. H. Pieper. 1995. From xenobiotic to antibiotic. Formation of protoanemonin from 4-chlorocatechol by enzymes of the 3-oxoadipate pathway. J. Biol. Chem. 27029229-29235. - PubMed
-
- Bradford, M. M. 1976. A rapid and sensitive method for the quantitation of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72248-254. - PubMed
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

