A critical assessment of the factors affecting reporter gene assays for promoter SNP function: a reassessment of -308 TNF polymorphism function using a novel integrated reporter system
- PMID: 19471307
- PMCID: PMC2986691
- DOI: 10.1038/ejhg.2009.80
A critical assessment of the factors affecting reporter gene assays for promoter SNP function: a reassessment of -308 TNF polymorphism function using a novel integrated reporter system
Abstract
One of the greatest challenges facing genetics is the development of strategies to identify functionally relevant genetic variation. The most common test of function is the reporter gene assay, in which allelic regulatory regions are used to drive the expression of a reporter gene, and differences in expression in a cell line after transient transfection are taken to be a reflection of the polymorphism. Many studies have reported small differences in single nucleotide polymorphism (SNP)-specific reporter activity, including the tumor necrosis factor (TNF) G-308A polymorphism. However, we have established that many variables inherent in the reporter gene approach can account for the reported allelic differences. Variables, such as the amount of DNA used in transfection, the amount of transfection control vector used, the method of transfection, the growth history of the host cells, and the quality and purity of DNA used, all influence TNF -308 SNP-specific transient reporter gene assays and serve as a caution for those researchers who apply this method to the functional assessment of polymorphic promoter sequences. We have developed an integrated reporter system that obviates some of these problems and shows that the TNF G-308A polymorphism is functionally relevant in this improved assay, thus confirming that the -308A allele expresses at a higher level compared with the -308G allele.
Figures
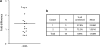
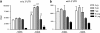
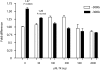


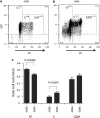
Similar articles
-
Relevance of the tumor necrosis factor alpha (TNF alpha) -308 promoter polymorphism in TNF alpha gene regulation.J Inflamm. 1995-1996;46(1):32-41. J Inflamm. 1995. PMID: 8832970
-
Interaction of the -308G/A promoter polymorphism of the tumor necrosis factor-alpha gene with single-nucleotide polymorphism 45 of the adiponectin gene: effect on serum adiponectin concentrations in a Spanish population.Clin Chem. 2006 Jan;52(1):97-103. doi: 10.1373/clinchem.2005.049452. Epub 2005 Oct 27. Clin Chem. 2006. PMID: 16254197
-
The -308 tumor necrosis factor-alpha promoter polymorphism effects transcription.Mol Immunol. 1997 Apr;34(5):391-9. doi: 10.1016/s0161-5890(97)00052-7. Mol Immunol. 1997. PMID: 9293772
-
Impact of the -308 TNF promoter polymorphism on the transcriptional regulation of the TNF gene: relevance to disease.J Leukoc Biol. 1999 Oct;66(4):562-6. doi: 10.1002/jlb.66.4.562. J Leukoc Biol. 1999. PMID: 10534109 Review.
-
Functional implications of genetic variation in non-coding DNA for disease susceptibility and gene regulation.Clin Sci (Lond). 2003 May;104(5):493-501. doi: 10.1042/CS20020304. Clin Sci (Lond). 2003. PMID: 12513691 Review.
Cited by
-
TNF-α, IL6, and IL10 polymorphisms and the effect of physical exercise on inflammatory parameters and physical performance in elderly women.Age (Dordr). 2013 Dec;35(6):2455-63. doi: 10.1007/s11357-013-9515-1. Epub 2013 Feb 21. Age (Dordr). 2013. PMID: 23430759 Free PMC article. Clinical Trial.
-
Association of a functional TNF variant with Plasmodium falciparum parasitaemia in a congolese population.Genes Immun. 2017 Sep;18(3):152-157. doi: 10.1038/gene.2017.13. Epub 2017 Jul 13. Genes Immun. 2017. PMID: 28703132
-
Molecular prioritization strategies to identify functional genetic variants in the cardiovascular disease-associated expression QTL Vanin-1.Eur J Hum Genet. 2014 May;22(5):688-95. doi: 10.1038/ejhg.2013.208. Epub 2013 Sep 18. Eur J Hum Genet. 2014. PMID: 24045843 Free PMC article.
-
Genetically determined high activities of the TNF-alpha, IL23/IL17, and NFkB pathways were associated with increased risk of ankylosing spondylitis.BMC Med Genet. 2018 Sep 12;19(1):165. doi: 10.1186/s12881-018-0680-z. BMC Med Genet. 2018. PMID: 30208882 Free PMC article.
-
Association between TNF promoter -308 G>A and LTA 252 A>G polymorphisms and systemic lupus erythematosus.Mol Biol Rep. 2014;41(4):2029-36. doi: 10.1007/s11033-014-3051-7. Epub 2014 Jan 14. Mol Biol Rep. 2014. PMID: 24420856
References
-
- Kraft P, Cox DG. Study designs for genome-wide association studies. Adv Genet. 2008;60:465–504. - PubMed
-
- Blangero J, Goring HH, Kent JW, Jr, et al. Quantitative trait nucleotide analysis using Bayesian model selection. Hum Biol. 2005;77:541–559. - PubMed
-
- Goring HH, Curran JE, Johnson MP, et al. Discovery of expression QTLs using large-scale transcriptional profiling in human lymphocytes. Nat Genet. 2007;39:1208–1216. - PubMed
-
- Buckland PR. The importance and identification of regulatory polymorphisms and their mechanisms of action. Biochim Biophys Acta. 2006;1762:17–28. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

