Modeling of solvent-dependent conformational transitions in Burkholderia cepacia lipase
- PMID: 19476626
- PMCID: PMC2695465
- DOI: 10.1186/1472-6807-9-38
Modeling of solvent-dependent conformational transitions in Burkholderia cepacia lipase
Abstract
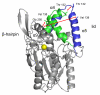
Background: The characteristic of most lipases is the interfacial activation at a lipid interface or in non-polar solvents. Interfacial activation is linked to a large conformational change of a lid, from a closed to an open conformation which makes the active site accessible for substrates. While for many lipases crystal structures of the closed and open conformation have been determined, the pathway of the conformational transition and possible bottlenecks are unknown. Therefore, molecular dynamics simulations of a closed homology model and an open crystal structure of Burkholderia cepacia lipase in water and toluene were performed to investigate the influence of solvents on structure, dynamics, and the conformational transition of the lid.
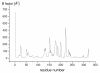
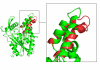
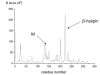
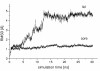
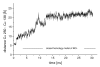
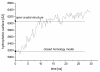
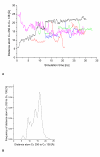
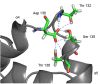
Results: The conformational transition of B. cepacia lipase was dependent on the solvent. In simulations of closed B. cepacia lipase in water no conformational transition was observed, while in three independent simulations of the closed lipase in toluene the lid gradually opened during the first 10-15 ns. The pathway of conformational transition was accessible and a barrier was identified, where a helix prevented the lid from opening to the completely open conformation. The open structure in toluene was stabilized by the formation of hydrogen bonds.In simulations of open lipase in water, the lid closed slowly during 30 ns nearly reaching its position in the closed crystal structure, while a further lid opening compared to the crystal structure was observed in toluene. While the helical structure of the lid was intact during opening in toluene, it partially unfolded upon closing in water. The closing of the lid in water was also observed, when with eight intermediate structures between the closed and the open conformation as derived from the simulations in toluene were taken as starting structures. A hydrophobic beta-hairpin was moving away from the lid in all simulations in water, which was not observed in simulations in toluene. The conformational transition of the lid was not correlated to the motions of the beta-hairpin structure.
Conclusion: Conformational transitions between the experimentally observed closed and open conformation of the lid were observed by multiple molecular dynamics simulations of B. cepacia lipase. Transitions in both directions occurred without applying restraints or external forces. The opening and closing were driven by the solvent and independent of a bound substrate molecule.
Figures
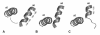
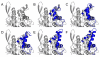
References
-
- Kazlauskas RJ, Weber HK. Improving hydrolases for organic synthesis. Curr Opin Chem Biol. 1998;2:121–126. - PubMed
-
- Bornscheuer UT, Kazlauskas RJ. Hydrolases in organic synthesis: regio- and stereolelective biotransformations. Weinheim: Wiley-VCH; 1999.
-
- Schmid RD, Verger R. Lipases: Interfacial enzymes with attractive applications. Angew Chem, Int Edit. 1998;37:1609–1633. - PubMed
-
- Jaeger KE, Eggert T. Lipases for biotechnology. Curr Opin Biotechnol. 2002;13:390–397. - PubMed
-
- Schrag JD, Cygler M. Lipases and alpha/beta hydrolase fold. Lipases, Part A. 1997;284:85–107. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

