Crystallization and preliminary crystallographic analysis of beta-L-arabinopyranosidase from Streptomyces avermitilis NBRC14893
- PMID: 19478450
- PMCID: PMC2688429
- DOI: 10.1107/S1744309109017230
Crystallization and preliminary crystallographic analysis of beta-L-arabinopyranosidase from Streptomyces avermitilis NBRC14893
Abstract
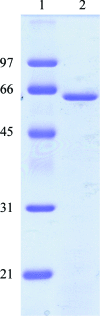
Beta-L-arabinopyranosidase from Streptomyces avermitilis NBRC14893 is a monomeric protein consisting of a catalytic domain belonging to glycosyl hydrolase family 27, an unknown domain and a substrate-binding domain belonging to carbohydrate-binding module family 13. The complete enzyme (residues 45-658) has successfully been cloned and homologously expressed in the Streptomyces expression system. beta-L-Arabinopyranosidase was crystallized by the sitting-drop vapour-diffusion method. The crystals diffracted to 1.6 A resolution and belonged to space group P2(1)2(1)2(1), with unit-cell parameters a = 68.2, b = 98.9, c = 181.3 A. The Matthews coefficient was calculated to be 2.38 A(3) Da(-1).
Figures

Similar articles
-
Crystallization of selenomethionyl exo-beta-1,3-galactanase from the basidiomycete Phanerochaete chrysosporium.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009 Dec 1;65(Pt 12):1274-6. doi: 10.1107/S1744309109043395. Epub 2009 Nov 27. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009. PMID: 20054127 Free PMC article.
-
A beta-l-Arabinopyranosidase from Streptomyces avermitilis is a novel member of glycoside hydrolase family 27.J Biol Chem. 2009 Sep 11;284(37):25097-106. doi: 10.1074/jbc.M109.022723. Epub 2009 Jul 16. J Biol Chem. 2009. PMID: 19608743 Free PMC article.
-
Crystallization and preliminary crystallographic analysis of exo-alpha-1,5-L-arabinofuranosidase from Streptomyces avermitilis NBRC14893.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2008 Nov 1;64(Pt 11):1007-9. doi: 10.1107/S1744309108029692. Epub 2008 Oct 25. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2008. PMID: 18997327 Free PMC article.
-
Crystallization and preliminary crystallographic analysis of the glycoside hydrolase family 115 α-glucuronidase from Streptomyces pristinaespiralis.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2011 Jan 1;67(Pt 1):68-71. doi: 10.1107/S1744309110043721. Epub 2010 Dec 22. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2011. PMID: 21206027 Free PMC article.
-
Structure and function of carbohydrate-binding module families 13 and 42 of glycoside hydrolases, comprising a β-trefoil fold.Biosci Biotechnol Biochem. 2013;77(7):1363-71. doi: 10.1271/bbb.130183. Epub 2013 Jul 7. Biosci Biotechnol Biochem. 2013. PMID: 23832347 Review.
Cited by
-
Characterization of an endo-processive-type xyloglucanase having a β-1,4-glucan-binding module and an endo-type xyloglucanase from Streptomyces avermitilis.Appl Environ Microbiol. 2012 Nov;78(22):7939-45. doi: 10.1128/AEM.01762-12. Epub 2012 Aug 31. Appl Environ Microbiol. 2012. PMID: 22941084 Free PMC article.
-
Arabinogalactan proteins: focus on carbohydrate active enzymes.Front Plant Sci. 2014 Jun 11;5:198. doi: 10.3389/fpls.2014.00198. eCollection 2014. Front Plant Sci. 2014. PMID: 24966860 Free PMC article. Review.
-
Crystallization of selenomethionyl exo-beta-1,3-galactanase from the basidiomycete Phanerochaete chrysosporium.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009 Dec 1;65(Pt 12):1274-6. doi: 10.1107/S1744309109043395. Epub 2009 Nov 27. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009. PMID: 20054127 Free PMC article.
-
A beta-l-Arabinopyranosidase from Streptomyces avermitilis is a novel member of glycoside hydrolase family 27.J Biol Chem. 2009 Sep 11;284(37):25097-106. doi: 10.1074/jbc.M109.022723. Epub 2009 Jul 16. J Biol Chem. 2009. PMID: 19608743 Free PMC article.
-
A comparison of gas stream cooling and plunge cooling of macromolecular crystals.J Appl Crystallogr. 2019 Aug 23;52(Pt 5):1222-1232. doi: 10.1107/S1600576719010318. eCollection 2019 Oct 1. J Appl Crystallogr. 2019. PMID: 31636524 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

