Fpocket: an open source platform for ligand pocket detection
- PMID: 19486540
- PMCID: PMC2700099
- DOI: 10.1186/1471-2105-10-168
Fpocket: an open source platform for ligand pocket detection
Abstract
Background: Virtual screening methods start to be well established as effective approaches to identify hits, candidates and leads for drug discovery research. Among those, structure based virtual screening (SBVS) approaches aim at docking collections of small compounds in the target structure to identify potent compounds. For SBVS, the identification of candidate pockets in protein structures is a key feature, and the recent years have seen increasing interest in developing methods for pocket and cavity detection on protein surfaces.
Results: Fpocket is an open source pocket detection package based on Voronoi tessellation and alpha spheres built on top of the publicly available package Qhull. The modular source code is organised around a central library of functions, a basis for three main programs: (i) Fpocket, to perform pocket identification, (ii) Tpocket, to organise pocket detection benchmarking on a set of known protein-ligand complexes, and (iii) Dpocket, to collect pocket descriptor values on a set of proteins. Fpocket is written in the C programming language, which makes it a platform well suited for the scientific community willing to develop new scoring functions and extract various pocket descriptors on a large scale level. Fpocket 1.0, relying on a simple scoring function, is able to detect 94% and 92% of the pockets within the best three ranked pockets from the holo and apo proteins respectively, outperforming the standards of the field, while being faster.
Conclusion: Fpocket provides a rapid, open source and stable basis for further developments related to protein pocket detection, efficient pocket descriptor extraction, or drugablity prediction purposes. Fpocket is freely available under the GNU GPL license at http://fpocket.sourceforge.net.
Figures
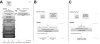
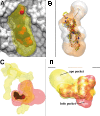
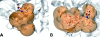
References
-
- Manly CJ, Chandrasekhar J, Ochterski JW, Hammer JD, Warfield BB. Strategies and tactics for optimizing the Hit-to-Lead process and beyond-A computational chemistry perspective. Drug Discov Today. 2008;13:99–109. - PubMed
-
- Villoutreix BO, Bastard K, Sperandio O, Fahraeus R, Poyet JL, Calvo F, Déprez B, Miteva MA. In silico-in vitro screening of protein-protein interactions: towards the next generation of therapeutics. Curr Pharm Biotechnol. 2008;9:103–22. - PubMed
-
- Levitt DG, Banaszak LJ. POCKET: a computer graphics method for identifying and displaying protein cavities and their surrounding amino acids. J Mol Graph. 1992;10:229–34. - PubMed
-
- Delaney JS. Finding and filling protein cavities using cellular logic operations. J Mol Graph. 1992;10:174–7. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

