Transcriptional regulation of the novel Toll-like receptor Tlr13
- PMID: 19487701
- PMCID: PMC2742818
- DOI: 10.1074/jbc.M109.022541
Transcriptional regulation of the novel Toll-like receptor Tlr13
Abstract
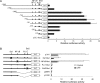
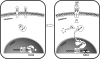
Little has been known about Tlr13 (Toll-like receptor 13), a novel member of the Toll-like receptor family. To elucidate the molecular basis of murine Tlr13 gene expression, the activity of the Tlr13 gene promoter was characterized. Reporter gene analysis and electrophoretic mobility shift assays demonstrated that Tlr13 gene transcription was regulated through three cis-acting elements that interacted with the Ets2, Sp1, and PU.1 transcription factors. Furthermore, our work suggests that these transcription factors may cooperate, culminating in maximal transcription of the Tlr13 gene. In contrast, NF-kappaB appeared to act as an inhibitor of Tlr13 transcription. Overexpression of Ets2 caused a strong increase in the transcriptional activity of the Tlr13 promoter; however, overexpression of NF-kappaB p65 dramatically inhibited it. Additionally, interferon-beta is capable of acting Tlr13 transcription, but the activated signaling of lipopolysaccharide/TLR4 and peptidoglycan/TLR2 strongly inhibited the Tlr13 gene promoter. Thus, these findings reveal the mechanism of Tlr13 gene regulation, thereby providing insight into the function of Tlr13 in the immune response to pathogen.
Figures



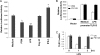
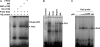

References
-
- Janeway C. A., Jr., Medzhitov R. (2002) Annu. Rev. Immunol. 20, 197–216 - PubMed
-
- Aderem A., Ulevitch R. J. (2000) Nature 406, 782–787 - PubMed
-
- Akira S., Uematsu S., Takeuchi O. (2006) Cell 124, 783–801 - PubMed
-
- Beutler B. (2002) Curr. Opin. Hematol. 9, 2–10 - PubMed
-
- Underhill D. M., Ozinsky A. (2002) Curr. Opin. Immunol. 14, 103–110 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

