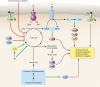
At the crossroads of bacterial metabolism and virulence factor synthesis in Staphylococci
- PMID: 19487727
- PMCID: PMC2698418
- DOI: 10.1128/MMBR.00005-09
At the crossroads of bacterial metabolism and virulence factor synthesis in Staphylococci
Abstract
Bacteria live in environments that are subject to rapid changes in the availability of the nutrients that are necessary to provide energy and biosynthetic intermediates for the synthesis of macromolecules. Consequently, bacterial survival depends on the ability of bacteria to regulate the expression of genes coding for enzymes required for growth in the altered environment. In pathogenic bacteria, adaptation to an altered environment often includes activating the transcription of virulence genes; hence, many virulence genes are regulated by environmental and nutritional signals. Consistent with this observation, the regulation of most, if not all, virulence determinants in staphylococci is mediated by environmental and nutritional signals. Some of these external signals can be directly transduced into a regulatory response by two-component regulators such as SrrAB; however, other external signals require transduction into intracellular signals. Many of the external environmental and nutritional signals that regulate virulence determinant expression can also alter bacterial metabolic status (e.g., iron limitation). Altering the metabolic status results in the transduction of external signals into intracellular metabolic signals that can be "sensed" by regulatory proteins (e.g., CodY, Rex, and GlnR). This review uses information derived primarily using Bacillus subtilis and Escherichia coli to articulate how gram-positive pathogens, with emphasis on Staphylococcus aureus and Staphylococcus epidermidis, regulate virulence determinant expression in response to a changing environment.
Figures
References
-
- Allard, M., H. Moisan, E. Brouillette, A. L. Gervais, M. Jacques, P. Lacasse, M. S. Diarra, and F. Malouin. 2006. Transcriptional modulation of some Staphylococcus aureus iron-regulated genes during growth in vitro and in a tissue cage model in vivo. Microbes Infect. 81679-1690. - PubMed
-
- Baba, T., F. Takeuchi, M. Kuroda, H. Yuzawa, K. Aoki, A. Oguchi, Y. Nagai, N. Iwama, K. Asano, T. Naimi, H. Kuroda, L. Cui, K. Yamamoto, and K. Hiramatsu. 2002. Genome and virulence determinants of high virulence community-acquired MRSA. Lancet 3591819-1827. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources