Analysis of Lsm1p and Lsm8p domains in the cellular localization of Lsm complexes in budding yeast
- PMID: 19490016
- PMCID: PMC2776932
- DOI: 10.1111/j.1742-4658.2009.07080.x
Analysis of Lsm1p and Lsm8p domains in the cellular localization of Lsm complexes in budding yeast
Abstract
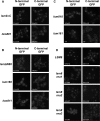
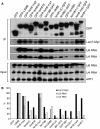
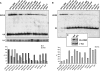
In eukaryotes, two heteroheptameric Sm-like (Lsm) complexes that differ by a single subunit localize to different cellular compartments and have distinct functions in RNA metabolism. The cytoplasmic Lsm1-7p complex promotes mRNA decapping and localizes to processing bodies, whereas the Lsm2-8p complex takes part in a variety of nuclear RNA processing events. The structural features that determine their different functions and localizations are not known. Here, we analyse a range of mutant and hybrid Lsm1 and Lsm8 proteins, shedding light on the relative importance of their various domains in determining their localization and ability to support growth. Although no single domain is either essential or sufficient for cellular localization, the Lsm1p N-terminus may act as part of a nuclear exclusion signal for Lsm1-7p, and the shorter Lsm8p N-terminus contributes to nuclear accumulation of Lsm2-8p. The C-terminal regions seem to play a secondary role in determining localization, with little or no contribution coming from the central Sm domains. The essential Lsm8 protein is remarkably resistant to mutation in terms of supporting viability, whereas Lsm1p appears more sensitive. These findings contribute to our understanding of how two very similar protein complexes can have different properties.
Figures






Similar articles
-
Molecular basis for the distinct cellular functions of the Lsm1-7 and Lsm2-8 complexes.RNA. 2020 Oct;26(10):1400-1413. doi: 10.1261/rna.075879.120. Epub 2020 Jun 9. RNA. 2020. PMID: 32518066 Free PMC article.
-
Requirements for nuclear localization of the Lsm2-8p complex and competition between nuclear and cytoplasmic Lsm complexes.J Cell Sci. 2007 Dec 15;120(Pt 24):4310-20. doi: 10.1242/jcs.019943. Epub 2007 Nov 20. J Cell Sci. 2007. PMID: 18029398 Free PMC article.
-
Nuclear pre-mRNA decapping and 5' degradation in yeast require the Lsm2-8p complex.Mol Cell Biol. 2004 Nov;24(21):9646-57. doi: 10.1128/MCB.24.21.9646-9657.2004. Mol Cell Biol. 2004. PMID: 15485930 Free PMC article.
-
Mutations in the Saccharomyces cerevisiae LSM1 gene that affect mRNA decapping and 3' end protection.Genetics. 2005 May;170(1):33-46. doi: 10.1534/genetics.104.034322. Epub 2005 Feb 16. Genetics. 2005. PMID: 15716506 Free PMC article.
-
Functions of Lsm proteins in mRNA degradation and splicing.Curr Opin Cell Biol. 2000 Jun;12(3):346-50. doi: 10.1016/s0955-0674(00)00098-3. Curr Opin Cell Biol. 2000. PMID: 10801455 Review.
Cited by
-
Prefoldins contribute to maintaining the levels of the spliceosome LSM2-8 complex through Hsp90 in Arabidopsis.Nucleic Acids Res. 2020 Jun 19;48(11):6280-6293. doi: 10.1093/nar/gkaa354. Nucleic Acids Res. 2020. PMID: 32396196 Free PMC article.
-
Molecular basis for the distinct cellular functions of the Lsm1-7 and Lsm2-8 complexes.RNA. 2020 Oct;26(10):1400-1413. doi: 10.1261/rna.075879.120. Epub 2020 Jun 9. RNA. 2020. PMID: 32518066 Free PMC article.
-
Malleable ribonucleoprotein machine: protein intrinsic disorder in the Saccharomyces cerevisiae spliceosome.PeerJ. 2013 Feb 12;1:e2. doi: 10.7717/peerj.2. Print 2013. PeerJ. 2013. PMID: 23638354 Free PMC article.
-
Both Sm-domain and C-terminal extension of Lsm1 are important for the RNA-binding activity of the Lsm1-7-Pat1 complex.RNA. 2012 May;18(5):936-44. doi: 10.1261/rna.029876.111. Epub 2012 Mar 26. RNA. 2012. PMID: 22450758 Free PMC article.
-
The life of U6 small nuclear RNA, from cradle to grave.RNA. 2018 Apr;24(4):437-460. doi: 10.1261/rna.065136.117. Epub 2018 Jan 24. RNA. 2018. PMID: 29367453 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

