Truncation of TRIM5 in the Feliformia explains the absence of retroviral restriction in cells of the domestic cat
- PMID: 19494015
- PMCID: PMC2715776
- DOI: 10.1128/JVI.00670-09
Truncation of TRIM5 in the Feliformia explains the absence of retroviral restriction in cells of the domestic cat
Abstract
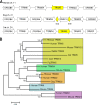
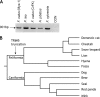
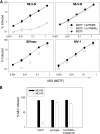
TRIM5alpha mediates a potent retroviral restriction phenotype in diverse mammalian species. Here, we identify a TRIM5 transcript in cat cells with a truncated B30.2 capsid binding domain and ablated restrictive function which, remarkably, is conserved across the Feliformia. Cat TRIM5 displayed no restriction activity, but ectopic expression conferred a dominant negative effect against human TRIM5alpha. Our findings explain the absence of retroviral restriction in cat cells and suggest that disruption of the TRIM5 locus has arisen independently at least twice in the Carnivora, with implications concerning the evolution of the host and pathogen in this taxon.
Figures
Similar articles
-
The B30.2(SPRY) domain of the retroviral restriction factor TRIM5alpha exhibits lineage-specific length and sequence variation in primates.J Virol. 2005 May;79(10):6111-21. doi: 10.1128/JVI.79.10.6111-6121.2005. J Virol. 2005. PMID: 15857996 Free PMC article.
-
All three variable regions of the TRIM5alpha B30.2 domain can contribute to the specificity of retrovirus restriction.J Virol. 2006 Sep;80(17):8554-65. doi: 10.1128/JVI.00688-06. J Virol. 2006. PMID: 16912305 Free PMC article.
-
An expanded clade of rodent Trim5 genes.Virology. 2009 Mar 15;385(2):473-83. doi: 10.1016/j.virol.2008.12.018. Epub 2009 Jan 15. Virology. 2009. PMID: 19147168 Free PMC article.
-
Control of viral infectivity by tripartite motif proteins.Hum Gene Ther. 2005 Oct;16(10):1125-32. doi: 10.1089/hum.2005.16.1125. Hum Gene Ther. 2005. PMID: 16218773 Free PMC article. Review.
-
The control of viral infection by tripartite motif proteins and cyclophilin A.Retrovirology. 2007 Jun 12;4:40. doi: 10.1186/1742-4690-4-40. Retrovirology. 2007. PMID: 17565686 Free PMC article. Review.
Cited by
-
Cellular restriction factors of feline immunodeficiency virus.Viruses. 2011 Oct;3(10):1986-2005. doi: 10.3390/v3101986. Epub 2011 Oct 21. Viruses. 2011. PMID: 22069525 Free PMC article. Review.
-
The fate of HIV-1 capsid: a biochemical assay for HIV-1 uncoating.Methods Mol Biol. 2014;1087:29-36. doi: 10.1007/978-1-62703-670-2_3. Methods Mol Biol. 2014. PMID: 24158811 Free PMC article.
-
Retroviral restriction and dependency factors in primates and carnivores.Vet Immunol Immunopathol. 2011 Oct 15;143(3-4):179-89. doi: 10.1016/j.vetimm.2011.06.002. Epub 2011 Jun 12. Vet Immunol Immunopathol. 2011. PMID: 21715018 Free PMC article. Review.
-
Prospects in Innate Immune Responses as Potential Control Strategies against Non-Primate Lentiviruses.Viruses. 2018 Aug 17;10(8):435. doi: 10.3390/v10080435. Viruses. 2018. PMID: 30126090 Free PMC article. Review.
-
Refrex-1, a soluble restriction factor against feline endogenous and exogenous retroviruses.J Virol. 2013 Nov;87(22):12029-40. doi: 10.1128/JVI.01267-13. Epub 2013 Aug 21. J Virol. 2013. PMID: 23966402 Free PMC article.
References
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215403-410. - PubMed
-
- Bainbridge, J. W., C. Stephens, K. Parsley, C. Demaison, A. Halfyard, A. J. Thrasher, and R. R. Ali. 2001. In vivo gene transfer to the mouse eye using an HIV-based lentiviral vector; efficient long-term transduction of corneal endothelium and retinal pigment epithelium. Gene Ther. 81665-1668. - PubMed
-
- Best, S., P. LeTissier, G. Towers, and J. P. Stoye. 1996. Positional cloning of the mouse retrovirus restriction gene Fv1. Nature 382826-829. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

