Evolution of alternative splicing regulation: changes in predicted exonic splicing regulators are not associated with changes in alternative splicing levels in primates
- PMID: 19495418
- PMCID: PMC2686173
- DOI: 10.1371/journal.pone.0005800
Evolution of alternative splicing regulation: changes in predicted exonic splicing regulators are not associated with changes in alternative splicing levels in primates
Abstract
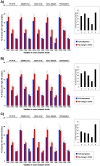
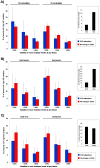
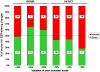
Alternative splicing is tightly regulated in a spatio-temporal and quantitative manner. This regulation is achieved by a complex interplay between spliceosomal (trans) factors that bind to different sequence (cis) elements. cis-elements reside in both introns and exons and may either enhance or silence splicing. Differential combinations of cis-elements allows for a huge diversity of overall splicing signals, together comprising a complex 'splicing code'. Many cis-elements have been identified, and their effects on exon inclusion levels demonstrated in reporter systems. However, the impact of interspecific differences in these elements on the evolution of alternative splicing levels has not yet been investigated at genomic level. Here we study the effect of interspecific differences in predicted exonic splicing regulators (ESRs) on exon inclusion levels in human and chimpanzee. For this purpose, we compiled and studied comprehensive datasets of predicted ESRs, identified by several computational and experimental approaches, as well as microarray data for changes in alternative splicing levels between human and chimpanzee. Surprisingly, we found no association between changes in predicted ESRs and changes in alternative splicing levels. This observation holds across different ESR exon positions, exon lengths, and 5' splice site strengths. We suggest that this lack of association is mainly due to the great importance of context for ESR functionality: many ESR-like motifs in primates may have little or no effect on splicing, and thus interspecific changes at short-time scales may primarily occur in these effectively neutral ESRs. These results underscore the difficulties of using current computational ESR prediction algorithms to identify truly functionally important motifs, and provide a cautionary tale for studies of the effect of SNPs on splicing in human disease.
Conflict of interest statement
Figures

Similar articles
-
Genome-wide prediction of cis-acting RNA elements regulating tissue-specific pre-mRNA alternative splicing.BMC Genomics. 2009 Jul 7;10 Suppl 1(Suppl 1):S4. doi: 10.1186/1471-2164-10-S1-S4. BMC Genomics. 2009. PMID: 19594881 Free PMC article.
-
Comparative analysis identifies exonic splicing regulatory sequences--The complex definition of enhancers and silencers.Mol Cell. 2006 Jun 23;22(6):769-781. doi: 10.1016/j.molcel.2006.05.008. Mol Cell. 2006. PMID: 16793546
-
Identification of conserved splicing motifs in mutually exclusive exons of 15 insect species.BMC Genomics. 2012 Apr 12;13 Suppl 2(Suppl 2):S1. doi: 10.1186/1471-2164-13-S2-S1. BMC Genomics. 2012. PMID: 22537296 Free PMC article.
-
Splicing regulation: from a parts list of regulatory elements to an integrated splicing code.RNA. 2008 May;14(5):802-13. doi: 10.1261/rna.876308. Epub 2008 Mar 27. RNA. 2008. PMID: 18369186 Free PMC article. Review.
-
[Molecular mechanism of mRNA alternative splicing].Yi Chuan Xue Bao. 2004 Jan;31(1):102-7. Yi Chuan Xue Bao. 2004. PMID: 15468927 Review. Chinese.
Cited by
-
The adaptive significance of unproductive alternative splicing in primates.RNA. 2010 Oct;16(10):2014-22. doi: 10.1261/rna.2127910. Epub 2010 Aug 18. RNA. 2010. PMID: 20719917 Free PMC article.
-
Origin and evolution of spliceosomal introns.Biol Direct. 2012 Apr 16;7:11. doi: 10.1186/1745-6150-7-11. Biol Direct. 2012. PMID: 22507701 Free PMC article. Review.
-
Functional properties and evolutionary splicing constraints on a composite exonic regulatory element of splicing in CFTR exon 12.Nucleic Acids Res. 2010 Jan;38(2):647-59. doi: 10.1093/nar/gkp1040. Epub 2009 Nov 12. Nucleic Acids Res. 2010. PMID: 19910374 Free PMC article.
-
Evolution of splicing regulatory networks in Drosophila.Genome Res. 2014 May;24(5):786-96. doi: 10.1101/gr.161521.113. Epub 2014 Feb 10. Genome Res. 2014. PMID: 24515119 Free PMC article.
-
Predicting functional alternative splicing by measuring RNA selection pressure from multigenome alignments.PLoS Comput Biol. 2009 Dec;5(12):e1000608. doi: 10.1371/journal.pcbi.1000608. Epub 2009 Dec 18. PLoS Comput Biol. 2009. PMID: 20019791 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

