A mathematical model to derive N-glycan structures and cellular enzyme activities from mass spectrometric data
- PMID: 19506293
- PMCID: PMC2757573
- DOI: 10.1093/glycob/cwp081
A mathematical model to derive N-glycan structures and cellular enzyme activities from mass spectrometric data
Abstract
Effective representation and characterization of biosynthetic pathways of glycosylation can be facilitated by mathematical modeling. This paper describes the expansion of a previously developed detailed model for N-linked glycosylation with the further application of the model to analyze MALDI-TOF mass spectra of human N-glycans in terms of underlying cellular enzyme activities. The glycosylation reaction network is automatically generated by the model, based on the reaction specificities of the glycosylation enzymes. The use of a molecular mass cutoff and a network pruning method typically limits the model size to about 10,000 glycan structures. This allows prediction of the complete glycan profile and its abundances for any set of assumed enzyme concentrations and reaction rate parameters. A synthetic mass spectrum from model-calculated glycan profiles is obtained and enzyme concentrations are adjusted to bring the theoretically calculated mass spectrum into agreement with experiment. The result of this process is a complete characterization of a measured glycan mass spectrum containing hundreds of masses in terms of the activities of 19 enzymes. In addition, a complete annotation of the mass spectrum in terms of glycan structure is produced, including the proportions of isomers within each peak. The method was applied to mass spectrometric data of normal human monocytes and monocytic leukemia (THP1) cells to derive glycosyltransferase activity changes underlying the differences in glycan structure between the normal and diseased cells. Model predictions could lead to a better understanding of the changes associated with disease states, identification of disease-associated biomarkers, and bioengineered glycan modifications.
Figures

 ), GN = N-acetylglucosamine (
), GN = N-acetylglucosamine ( ), G = glucose (
), G = glucose ( ).
).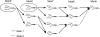

 ) and THP1 monocytic leukemia cells (
) and THP1 monocytic leukemia cells ( ) with synthetic mass spectrum peak areas calculated from the model. The values plotted are the relative peak areas as a percentage of the total area of the spectrum. Mass numbers from 1400 to 4000 were included. The raw data files were downloaded from the CFG website and processed with software developed as described in Material and methods. (http://www. functionalglycomics.org/glycomics/publicdata/glycoprofiling.jsp).
) with synthetic mass spectrum peak areas calculated from the model. The values plotted are the relative peak areas as a percentage of the total area of the spectrum. Mass numbers from 1400 to 4000 were included. The raw data files were downloaded from the CFG website and processed with software developed as described in Material and methods. (http://www. functionalglycomics.org/glycomics/publicdata/glycoprofiling.jsp).
 mannose,
mannose,  fucose,
fucose,  galactose,
galactose,  N-acetylglucosamine,
N-acetylglucosamine,  N-acetylgalactosamine,
N-acetylgalactosamine,  N-acetylneuraminic acid,
N-acetylneuraminic acid,  calculated,
calculated,  measured.
measured.
 ), F = fucose (
), F = fucose ( ), GN = N-acetylglucosamine (
), GN = N-acetylglucosamine ( ), M = mannose (
), M = mannose ( ), NN = N-acetylneuraminic acid (
), NN = N-acetylneuraminic acid ( ).
).References
-
- Banin E, Neuberger Y, Altshuler Y, Halevi A, Inbar O, Nir D, Dukler A. A novel linear code (R) nomenclature for complex carbohydrates. Trends Glycosci Glycotechnol. 2002;14:127–137.
-
- Bibila TA, Flickinger MC. A model of interorganelle monoclonal antibody transport and secretion in mouse hybridoma cells. Biotechnol Bioeng. 1991;38:767–780. - PubMed
-
- Brockhausen I, Kuhns W, Schachter H, Matta KL, Sutherland DR, Baker MA. Biosynthesis of O-glycans in leukocytes from normal donors and from patients with leukemia: Increase in O-glycan core 2 UDP-GlcNAc:Galb3GalNAca-R (GlcNAc to GalNAc) b(1-6)-N-acetylglucosaminyltransferase in leukemic cells. Cancer Res. 1991;51:1257–1263. - PubMed
-
- Brockhausen I, Schutzbach J, Kuhns W. Glycoproteins and their relationship to human disease. Acta Anat. 1998;161:36–78. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

