Poly(A) tail length is controlled by the nuclear poly(A)-binding protein regulating the interaction between poly(A) polymerase and the cleavage and polyadenylation specificity factor
- PMID: 19509282
- PMCID: PMC2755688
- DOI: 10.1074/jbc.M109.018226
Poly(A) tail length is controlled by the nuclear poly(A)-binding protein regulating the interaction between poly(A) polymerase and the cleavage and polyadenylation specificity factor
Abstract
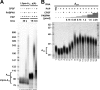
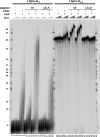
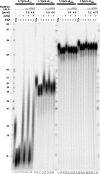
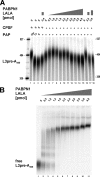
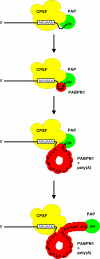
Poly(A) tails of mRNAs are synthesized in the cell nucleus with a defined length, approximately 250 nucleotides in mammalian cells. The same type of length control is seen in an in vitro polyadenylation system reconstituted from three proteins: poly(A) polymerase, cleavage and polyadenylation specificity factor (CPSF), and the nuclear poly(A)-binding protein (PABPN1). CPSF, binding the polyadenylation signal AAUAAA, and PABPN1, binding the growing poly(A) tail, cooperatively stimulate poly(A) polymerase such that a complete poly(A) tail is synthesized in one processive event, which terminates at a length of approximately 250 nucleotides. We report that PABPN1 is required to restrict CPSF binding to the AAUAAA sequence and to permit the stimulation of poly(A) polymerase by AAUAAA-bound CPSF to be maintained throughout the elongation reaction. The stimulation by CPSF is disrupted when the poly(A) tail has reached a length of approximately 250 nucleotides, and this terminates processive elongation. PABPN1 measures the length of the tail and is responsible for disrupting the CPSF-poly(A) polymerase interaction.
Figures



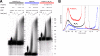
References
-
- Meyer S., Temme C., Wahle E. (2004) Crit. Rev. Biochem. Mol. Biol. 39,197–216 - PubMed
-
- Parker R., Song H. (2004) Nat. Struct. Mol. Biol. 11,121–127 - PubMed
-
- Wilson T., Treisman R. (1988) Nature 336,396–399 - PubMed
-
- Sachs A. (2000) in Translational Control of Gene Expression ( Sonenberg N., Hershey J. W. B., Mathews M. B. eds) pp. 447–465, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

