Analysis of DNA relaxation and cleavage activities of recombinant Mycobacterium tuberculosis DNA topoisomerase I from a new expression and purification protocol
- PMID: 19519900
- PMCID: PMC2702276
- DOI: 10.1186/1471-2091-10-18
Analysis of DNA relaxation and cleavage activities of recombinant Mycobacterium tuberculosis DNA topoisomerase I from a new expression and purification protocol
Abstract
Background: Mycobacterium tuberculosis DNA topoisomerase I is an attractive target for discovery of novel TB drugs that act by enhancing the accumulation of the topoisomerase-DNA cleavage product. It shares a common transesterification domain with other type IA DNA topoisomerases. There is, however, no homology between the C-terminal DNA binding domains of Escherichia coli and M. tuberculosis DNA topoisomerase I proteins.
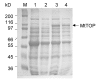
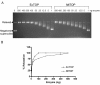
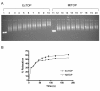
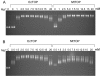
Results: A new protocol for expression and purification of recombinant M. tuberculosis DNA topoisomerase I (MtTOP) has been developed to produce enzyme of much higher specific activity than previously characterized recombinant enzyme. MtTOP was found to be less efficient than E. coli DNA topoisomerase I (EcTOP) in removal of remaining negative supercoils from partially relaxed DNA. DNA cleavage by MtTOP was characterized for the first time. Comparison of DNA cleavage site selectivity with EcTOP showed differences in cleavage site preferences, but the preferred sites of both enzymes have a C nucleotide in the -4 position.
Conclusion: Recombinant M. tuberculosis DNA topoisomerase I can be expressed as a soluble protein and purified in high yield from E. coli host with a new protocol. Analysis of DNA cleavage with M. tuberculosis DNA substrate showed that the preferred DNA cleavage sites have a C nucleotide in the -4 position.
Figures


Similar articles
-
The DNA relaxation activity and covalent complex accumulation of Mycobacterium tuberculosis topoisomerase I can be assayed in Escherichia coli: application for identification of potential FRET-dye labeling sites.BMC Biochem. 2010 Sep 30;11:41. doi: 10.1186/1471-2091-11-41. BMC Biochem. 2010. PMID: 20920291 Free PMC article.
-
Characterization of DNA topoisomerase I from Mycobacterium tuberculosis: DNA cleavage and religation properties and inhibition of its activity.Arch Biochem Biophys. 2012 Dec 15;528(2):197-203. doi: 10.1016/j.abb.2012.10.004. Epub 2012 Oct 17. Arch Biochem Biophys. 2012. PMID: 23085346
-
Cloning, expression, purification and characterization of DNA topoisomerase I of Mycobacterium tuberculosis.Gene. 1996 Oct 31;178(1-2):63-9. doi: 10.1016/0378-1119(96)00335-6. Gene. 1996. PMID: 8921893
-
DNA topoisomerase I and DNA gyrase as targets for TB therapy.Drug Discov Today. 2017 Mar;22(3):510-518. doi: 10.1016/j.drudis.2016.11.006. Epub 2016 Nov 14. Drug Discov Today. 2017. PMID: 27856347 Review.
-
DNA topoisomerase I from mycobacteria--a potential drug target.Curr Pharm Des. 2002;8(22):1995-2007. doi: 10.2174/1381612023393567. Curr Pharm Des. 2002. PMID: 12171525 Review.
Cited by
-
Metal ion and inter-domain interactions as functional networks in E. coli topoisomerase I.Gene. 2013 Jul 25;524(2):253-60. doi: 10.1016/j.gene.2013.04.008. Epub 2013 Apr 20. Gene. 2013. PMID: 23612251 Free PMC article.
-
Identification of anziaic acid, a lichen depside from Hypotrachyna sp., as a new topoisomerase poison inhibitor.PLoS One. 2013 Apr 8;8(4):e60770. doi: 10.1371/journal.pone.0060770. Print 2013. PLoS One. 2013. PMID: 23593306 Free PMC article.
-
The interaction between transport-segment DNA and topoisomerase IA-crystal structure of MtbTOP1 in complex with both G- and T-segments.Nucleic Acids Res. 2023 Jan 11;51(1):349-364. doi: 10.1093/nar/gkac1205. Nucleic Acids Res. 2023. PMID: 36583363 Free PMC article.
-
Localization of Mycobacterium tuberculosis topoisomerase I C-terminal sequence motif required for inhibition by endogenous toxin MazF4.Front Microbiol. 2022 Dec 5;13:1032320. doi: 10.3389/fmicb.2022.1032320. eCollection 2022. Front Microbiol. 2022. PMID: 36545199 Free PMC article.
-
Mechanism of Type IA Topoisomerases.Molecules. 2020 Oct 17;25(20):4769. doi: 10.3390/molecules25204769. Molecules. 2020. PMID: 33080770 Free PMC article. Review.
References
-
- Corbett KD, Berger JM. Structure, molecular mechanisms, and evolutionary relationships in DNA topoisomerases. Annu Rev Biophys Biomol Struct. 2004;33:95–118. - PubMed
-
- Wang JC. Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol. 2002;3:430–440. - PubMed
-
- Tse-Dinh Y- Exploring DNA Topoisomerases as Targets of Novel Therapeutic Agents in the Treatment of Infectious Diseases. Infectious Disorders – Drug Targets. 2007;7:3–9. - PubMed
-
- Drlica K. Mechanism of fluoroquinolone action. Curr Opin Microbiol. 1999;2:504–508. - PubMed
-
- Hooper DC. Bacterial topoisomerases, anti-topoisomerases, and anti-topoisomerase resistance. Clin Infect Dis. 1998;27:S54–63. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

