Detection of single nucleotide variations in expressed exons of the human genome using RNA-Seq
- PMID: 19528076
- PMCID: PMC2760790
- DOI: 10.1093/nar/gkp507
Detection of single nucleotide variations in expressed exons of the human genome using RNA-Seq
Abstract
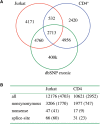
Whole-genome resequencing is still a costly method to detect genetic mutations that lead to altered forms of proteins and may be associated with disease development. Since the majority of disease-related single nucleotide variations (SNVs) are found in protein-coding regions, we propose to identify SNVs in expressed exons of the human genome using the recently developed RNA-Seq technique. We identify 12 176 and 10 621 SNVs, respectively, in Jurkat T cells and CD4(+) T cells from a healthy donor. Interestingly, our data show that one copy of the TAL-1 proto-oncogene has a point mutation in 3' UTR and only the mutant allele is expressed in Jurkat cells. We provide a comprehensive dataset for further understanding the cancer biology of Jurkat cells. Our results indicate that this is a cost-effective and efficient strategy to systematically identify SNVs in the expressed regions of the human genome.
Figures

References
-
- Stenson PD, Ball EV, Mort M, Phillips AD, Shiel JA, Thomas NS, Abeysinghe S, Krawczak M, Cooper DN. Human Gene Mutation Database (HGMD): 2003 update. Hum. Mutat. 2003;21:577–581. - PubMed
-
- Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods. 2008;5:621–628. - PubMed
-
- Sultan M, Schulz MH, Richard H, Magen A, Klingenhoff A, Scherf M, Seifert M, Borodina T, Soldatov A, Parkhomchuk D, et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science. 2008;321:956–960. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

