Comments on the analysis of unbalanced microarray data
- PMID: 19528084
- PMCID: PMC2732368
- DOI: 10.1093/bioinformatics/btp363
Comments on the analysis of unbalanced microarray data
Abstract
Motivation: Permutation testing is very popular for analyzing microarray data to identify differentially expressed (DE) genes; estimating false discovery rates (FDRs) is a very popular way to address the inherent multiple testing problem. However, combining these approaches may be problematic when sample sizes are unequal.
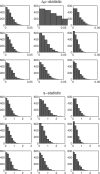
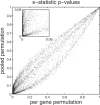
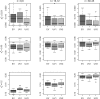
Results: With unbalanced data, permutation tests may not be suitable because they do not test the hypothesis of interest. In addition, permutation tests can be biased. Using biased P-values to estimate the FDR can produce unacceptable bias in those estimates. Results also show that the approach of pooling permutation null distributions across genes can produce invalid P-values, since even non-DE genes can have different permutation null distributions. We encourage researchers to use statistics that have been shown to reliably discriminate DE genes, but caution that associated P-values may be either invalid, or a less-effective metric for discriminating DE genes.
Figures


References
-
- Allison DB, et al. A mixture model approach for the analysis of microarray gene expression data. Comput. Stat. Data Anal. 2002;39:1–20.
-
- Allison DB, et al. Microarray data analysis: from disarray to consolidation and consensus. Nat. Rev. Genet. 2006;7:55–65. - PubMed
-
- Benjamini Y, Hochberg Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Methodol. 1995;57:289–300.
-
- Calian V, et al. Partitioning to uncover conditions for permutation tests to control multiple testing error rates. Biom. J. 2008;50:756–766. - PubMed
-
- Cheng C, et al. Statistical significance threshold criteria for analysis of microarray gene expression data. Stat. Appl. Genet. Mol. Biol. 2004;3:36. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

