IGF2BP1 enhances HCV IRES-mediated translation initiation via the 3'UTR
- PMID: 19541769
- PMCID: PMC2714754
- DOI: 10.1261/rna.1578409
IGF2BP1 enhances HCV IRES-mediated translation initiation via the 3'UTR
Abstract
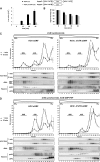
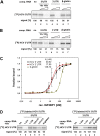
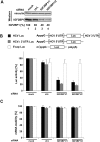
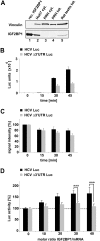
The positive-strand RNA genome of the Hepatitis C virus (HCV) contains an internal ribosome entry site (IRES) in the 5'untranslated region (5'UTR) and structured sequence elements within the 3'UTR, but no poly(A) tail. Employing a limited set of initiation factors, the HCV IRES coordinates the 5'cap-independent assembly of the 43S pre-initiation complex at an internal initiation codon located in the IRES sequence. We have established a Huh7 cell-derived in vitro translation system that shows a 3'UTR-dependent enhancement of 43S pre-initiation complex formation at the HCV IRES. Through the use of tobramycin (Tob)-aptamer affinity chromatography, we identified the Insulin-like growth factor-II mRNA-binding protein 1 (IGF2BP1) as a factor that interacts with both, the HCV 5'UTR and 3'UTR. We report that IGF2BP1 specifically enhances translation at the HCV IRES, but it does not affect 5'cap-dependent translation. RNA interference against IGF2BP1 in HCV replicon RNA-containing Huh7 cells reduces HCV IRES-mediated translation, whereas replication remains unaffected. Interestingly, we found that endogenous IGF2BP1 specifically co-immunoprecipitates with HCV replicon RNA, the ribosomal 40S subunit, and eIF3. Furthermore eIF3 comigrates with IGF2BP1 in 80S ribosomal complexes when a reporter mRNA bearing both the HCV 5'UTR and HCV 3'UTR is translated. Our data suggest that IGF2BP1, by binding to the HCV 5'UTR and/or HCV 3'UTR, recruits eIF3 and enhances HCV IRES-mediated translation.
Figures

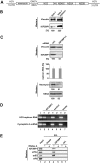

Similar articles
-
Hepatitis C Virus Translation Regulation.Int J Mol Sci. 2020 Mar 27;21(7):2328. doi: 10.3390/ijms21072328. Int J Mol Sci. 2020. PMID: 32230899 Free PMC article. Review.
-
Poly(C)-binding protein 2 interacts with sequences required for viral replication in the hepatitis C virus (HCV) 5' untranslated region and directs HCV RNA replication through circularizing the viral genome.J Virol. 2011 Aug;85(16):7954-64. doi: 10.1128/JVI.00339-11. Epub 2011 Jun 1. J Virol. 2011. PMID: 21632751 Free PMC article.
-
Hepatitis-C-virus-like internal ribosome entry sites displace eIF3 to gain access to the 40S subunit.Nature. 2013 Nov 28;503(7477):539-43. doi: 10.1038/nature12658. Epub 2013 Nov 3. Nature. 2013. PMID: 24185006 Free PMC article.
-
Unlike for cellular mRNAs and other viral internal ribosome entry sites (IRESs), the eIF3 subunit e is not required for the translational activity of the HCV IRES.J Biol Chem. 2020 Feb 14;295(7):1843-1856. doi: 10.1074/jbc.RA119.009502. Epub 2020 Jan 12. J Biol Chem. 2020. PMID: 31929110 Free PMC article.
-
Structural and mechanistic insights into hepatitis C viral translation initiation.Nat Rev Microbiol. 2007 Jan;5(1):29-38. doi: 10.1038/nrmicro1558. Epub 2006 Nov 27. Nat Rev Microbiol. 2007. PMID: 17128284 Review.
Cited by
-
In-cell SHAPE uncovers dynamic interactions between the untranslated regions of the foot-and-mouth disease virus RNA.Nucleic Acids Res. 2017 Feb 17;45(3):1416-1432. doi: 10.1093/nar/gkw795. Nucleic Acids Res. 2017. PMID: 28180318 Free PMC article.
-
Lipid Droplet Isolation for Quantitative Mass Spectrometry Analysis.J Vis Exp. 2017 Apr 17;(122):55585. doi: 10.3791/55585. J Vis Exp. 2017. PMID: 28448054 Free PMC article.
-
RNA structural elements of hepatitis C virus controlling viral RNA translation and the implications for viral pathogenesis.Viruses. 2012 Oct 19;4(10):2233-50. doi: 10.3390/v4102233. Viruses. 2012. PMID: 23202462 Free PMC article. Review.
-
Alternative Mechanisms to Initiate Translation in Eukaryotic mRNAs.Comp Funct Genomics. 2012;2012:391546. doi: 10.1155/2012/391546. Epub 2012 Feb 16. Comp Funct Genomics. 2012. PMID: 22536116 Free PMC article.
-
IGF2BP1-An Oncofetal RNA-Binding Protein Fuels Tumor Virus Propagation.Viruses. 2023 Jun 24;15(7):1431. doi: 10.3390/v15071431. Viruses. 2023. PMID: 37515119 Free PMC article. Review.
References
-
- Anthony D, Merrick WC. Analysis of 40 S and 80 S complexes with mRNA as measured by sucrose density gradients and primer extension inhibition. J Biol Chem. 1992;267:1554–1562. - PubMed
-
- Atlas R, Behar L, Sapoznik S, Ginzburg I. Dynamic association with polysomes during P19 neuronal differentiation and an untranslated-region-dependent translation regulation of the τ mRNA by the τ mRNA-associated proteins IMP1, HuD, and G3BP1. J Neurosci Res. 2007;85:173–183. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
