CtrA, a global response regulator, uses a distinct second category of weak DNA binding sites for cell cycle transcription control in Caulobacter crescentus
- PMID: 19542275
- PMCID: PMC2725627
- DOI: 10.1128/JB.00355-09
CtrA, a global response regulator, uses a distinct second category of weak DNA binding sites for cell cycle transcription control in Caulobacter crescentus
Abstract
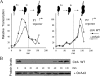
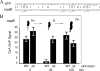
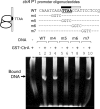
CtrA controls cell cycle programs of chromosome replication and genetic transcription. Phosphorylated CtrA approximately P exhibits high affinity (dissociation constant [K(d)], <10 nM) for consensus TTAA-N7-TTAA binding sites with "typical" (N = 7) spacing. We show here that ctrA promoters P1 and P2 use low-affinity (K(d), >500 nM) CtrA binding sites with "atypical" (N not equal 7) spacing. Footprints demonstrated that phosphorylated CtrA approximately P does not exhibit increased affinity for "atypical" sites, as it does for sites in the replication origin. Instead, high levels of CtrA (>10 microM) accumulate, which can drive CtrA binding to "atypical" sites. In vivo cross-linking showed that when the stable CtrADelta3 protein persists during the cell cycle, the "atypical" sites at ctrA and motB are persistently bound. Interestingly, the cell cycle timing of ctrA P1 and P2 transcription is not altered by persistent CtrADelta3 binding. Therefore, operator DNA occupancy is not sufficient for regulation, and it is the cell cycle variation of CtrA approximately P phosphorylation that provides the dominant "activation" signal. Protein dimerization is one potential means of "activation." The glutathione S-transferase (GST) protein dimerizes, and fusion with CtrA (GST-CtrA) creates a stable dimer with enhanced affinity for TTAA motifs. Electrophoretic mobility shift assays with GST-CtrA revealed cooperative modes of binding that further distinguish the "atypical" sites. GST-CtrA also binds a single TTAA motif in ctrA P1 aided by DNA in the extended TTAACCAT motif. We discuss how "atypical" sites are a common yet distinct category of CtrA regulatory sites and new implications for the working and evolution of cell cycle control networks.
Figures


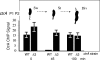



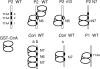
Similar articles
-
Analysis of a cell-cycle promoter bound by a response regulator.J Mol Biol. 2000 Sep 29;302(4):761-75. doi: 10.1006/jmbi.2000.4500. J Mol Biol. 2000. PMID: 10993722
-
Feedback control of a master bacterial cell-cycle regulator.Proc Natl Acad Sci U S A. 1999 Jun 8;96(12):6648-53. doi: 10.1073/pnas.96.12.6648. Proc Natl Acad Sci U S A. 1999. PMID: 10359766 Free PMC article.
-
Cell cycle regulator phosphorylation stimulates two distinct modes of binding at a chromosome replication origin.EMBO J. 2000 Mar 1;19(5):1138-47. doi: 10.1093/emboj/19.5.1138. EMBO J. 2000. PMID: 10698954 Free PMC article.
-
A genetic oscillator and the regulation of cell cycle progression in Caulobacter crescentus.Cell Cycle. 2004 Oct;3(10):1252-4. doi: 10.4161/cc.3.10.1181. Epub 2004 Oct 17. Cell Cycle. 2004. PMID: 15467452 Review.
-
Who's in charge here? Regulating cell cycle regulators.Curr Opin Microbiol. 2008 Dec;11(6):547-52. doi: 10.1016/j.mib.2008.09.019. Epub 2008 Nov 3. Curr Opin Microbiol. 2008. PMID: 18955157 Review.
Cited by
-
The Caulobacter crescentus ctrA P1 promoter is essential for the coordination of cell cycle events that prevent the overinitiation of DNA replication.Microbiology (Reading). 2012 Oct;158(Pt 10):2492-2503. doi: 10.1099/mic.0.055285-0. Epub 2012 Jul 12. Microbiology (Reading). 2012. PMID: 22790399 Free PMC article.
-
Periplasmic protein EipA determines envelope stress resistance and virulence in Brucella abortus.Mol Microbiol. 2019 Mar;111(3):637-661. doi: 10.1111/mmi.14178. Epub 2019 Jan 21. Mol Microbiol. 2019. PMID: 30536925 Free PMC article.
-
Loss of the response regulator CtrA causes pleiotropic effects on gene expression but does not affect growth phase regulation in Rhodobacter capsulatus.J Bacteriol. 2010 Jun;192(11):2701-10. doi: 10.1128/JB.00160-10. Epub 2010 Apr 2. J Bacteriol. 2010. PMID: 20363938 Free PMC article.
-
Transcriptional regulation of the Rhodobacter capsulatus response regulator CtrA.Microbiology (Reading). 2013 Jan;159(Pt 1):96-106. doi: 10.1099/mic.0.062349-0. Epub 2012 Nov 15. Microbiology (Reading). 2013. PMID: 23154973 Free PMC article.
-
Getting in the loop: regulation of development in Caulobacter crescentus.Microbiol Mol Biol Rev. 2010 Mar;74(1):13-41. doi: 10.1128/MMBR.00040-09. Microbiol Mol Biol Rev. 2010. PMID: 20197497 Free PMC article. Review.
References
-
- Bellefontaine, A.-F., C. E. Pierreux, P. Mertens, J. Vandenhaute, J.-J. Letesson, and X. De Bolle. 2002. Plasticity of a transcriptional regulatory network among alpha-proteobacteria is supported by the identification of CtrA targets in Brucella abortus. Mol. Microbiol. 43945-960. - PubMed
-
- Biondi, E. G., S. J. Reisinger, J. M. Skerker, M. Arif, B. S. Perchuk, K. R. Ryan, and M. T. Laub. 2006. Regulation of the bacterial cell cycle by an integrated genetic circuit. Nature 444899-904. - PubMed
-
- Domian, I. J., K. C. Quon, and L. Shapiro. 1997. Cell type-specific phosphorylation and proteolysis of a transcriptional regulator controls the G1 to S transition in a bacterial cell cycle. Cell 90415-424. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

