Proteomic and genomic characterization of highly infectious Clostridium difficile 630 spores
- PMID: 19542279
- PMCID: PMC2725610
- DOI: 10.1128/JB.00597-09
Proteomic and genomic characterization of highly infectious Clostridium difficile 630 spores
Abstract
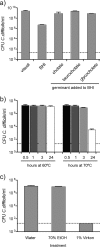
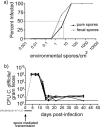
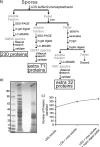
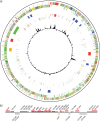
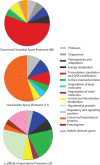
Clostridium difficile, a major cause of antibiotic-associated diarrhea, produces highly resistant spores that contaminate hospital environments and facilitate efficient disease transmission. We purified C. difficile spores using a novel method and show that they exhibit significant resistance to harsh physical or chemical treatments and are also highly infectious, with <7 environmental spores per cm(2) reproducibly establishing a persistent infection in exposed mice. Mass spectrometric analysis identified approximately 336 spore-associated polypeptides, with a significant proportion linked to translation, sporulation/germination, and protein stabilization/degradation. In addition, proteins from several distinct metabolic pathways associated with energy production were identified. Comparison of the C. difficile spore proteome to those of other clostridial species defined 88 proteins as the clostridial spore "core" and 29 proteins as C. difficile spore specific, including proteins that could contribute to spore-host interactions. Thus, our results provide the first molecular definition of C. difficile spores, opening up new opportunities for the development of diagnostic and therapeutic approaches.
Figures

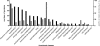
References
-
- Allen, J. E., M. C. Cerrone, P. R. Beatty, and R. S. Stephens. 1990. Cysteine-rich outer membrane proteins of Chlamydia trachomatis display compensatory sequence changes between biovariants. Mol. Microbiol. 41543-1550. - PubMed
-
- Bartlett, J. G. 2006. The new epidemic of Clostridium difficile-associated enteric disease. Ann. Intern. Med. 145758-764. - PubMed
-
- Bartlett, J. G., T. Chang, N. S. Taylor, and A. B. Onderdonk. 1979. Colitis induced by Clostridium difficile. Rev. Infect. Dis. 1370-378. - PubMed
-
- Borriello, S. P. 1998. Pathogenesis of Clostridium difficile infection. J. Antimicrob. Chemother. 41(Suppl. C)13-19. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

