Effect of perturbation of ATP level on the activity and regulation of nitrogenase in Rhodospirillum rubrum
- PMID: 19542280
- PMCID: PMC2725632
- DOI: 10.1128/JB.00585-09
Effect of perturbation of ATP level on the activity and regulation of nitrogenase in Rhodospirillum rubrum
Abstract
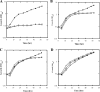
Nitrogenase activity in Rhodospirillum rubrum and in some other photosynthetic bacteria is regulated in part by the availability of light. This regulation is through a posttranslational modification system that is itself regulated by P(II) homologs in the cell. P(II) is one of the most broadly distributed regulatory proteins in nature and directly or indirectly senses nitrogen and carbon signals in the cell. However, its possible role in responding to light availability remains unclear. Because P(II) binds ATP, we tested the hypothesis that removal of light would affect P(II) by changing intracellular ATP levels, and this in turn would affect the regulation of nitrogenase activity. This in vivo test involved a variety of different methods for the measurement of ATP, as well as the deliberate perturbation of intracellular ATP levels by chemical and genetic means. To our surprise, we found fairly normal levels of nitrogenase activity and posttranslational regulation of nitrogenase even under conditions of drastically reduced ATP levels. This indicates that low ATP levels have no more than a modest impact on the P(II)-mediated regulation of NifA activity and on the posttranslational regulation of nitrogenase activity. The relatively high nitrogenase activity also shows that the ATP-dependent electron flux from dinitrogenase reductase to dinitrogenase is also surprisingly insensitive to a depleted ATP level. These in vivo results disprove the simple model of ATP as the key energy signal to P(II) under these conditions. We currently suppose that the ratio of ADP/ATP might be the relevant signal, as suggested by a number of recent in vitro analyses.
Figures



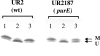
Similar articles
-
Effect of AmtB homologues on the post-translational regulation of nitrogenase activity in response to ammonium and energy signals in Rhodospirillum rubrum.Microbiology (Reading). 2006 Jul;152(Pt 7):2075-2089. doi: 10.1099/mic.0.28903-0. Microbiology (Reading). 2006. PMID: 16804182
-
Mutagenesis and functional characterization of the glnB, glnA, and nifA genes from the photosynthetic bacterium Rhodospirillum rubrum.J Bacteriol. 2000 Feb;182(4):983-92. doi: 10.1128/JB.182.4.983-992.2000. J Bacteriol. 2000. PMID: 10648524 Free PMC article.
-
GlnD is essential for NifA activation, NtrB/NtrC-regulated gene expression, and posttranslational regulation of nitrogenase activity in the photosynthetic, nitrogen-fixing bacterium Rhodospirillum rubrum.J Bacteriol. 2005 Feb;187(4):1254-65. doi: 10.1128/JB.187.4.1254-1265.2005. J Bacteriol. 2005. PMID: 15687189 Free PMC article.
-
H2 metabolism in photosynthetic bacteria and relationship to N2 fixation.Ann Microbiol (Paris). 1983 Jul-Aug;134B(1):115-35. doi: 10.1016/s0769-2609(83)80100-8. Ann Microbiol (Paris). 1983. PMID: 6139053 Review.
-
Metabolic regulation of nitrogen fixation in Rhodospirillum rubrum.Biochem Soc Trans. 2006 Feb;34(Pt 1):160-1. doi: 10.1042/BST0340160. Biochem Soc Trans. 2006. PMID: 16417510 Review.
Cited by
-
The poor growth of Rhodospirillum rubrum mutants lacking RubisCO is due to the accumulation of ribulose-1,5-bisphosphate.J Bacteriol. 2011 Jul;193(13):3293-303. doi: 10.1128/JB.00265-11. Epub 2011 Apr 29. J Bacteriol. 2011. PMID: 21531802 Free PMC article.
-
Elimination of Rubisco alters the regulation of nitrogenase activity and increases hydrogen production in Rhodospirillum rubrum.Int J Hydrogen Energy. 2010 Jul 1;35(14):7377-7385. doi: 10.1016/j.ijhydene.2010.04.183. Int J Hydrogen Energy. 2010. PMID: 20652089 Free PMC article.
-
Transcriptomics reveal several gene expression patterns in the piezophile Desulfovibrio hydrothermalis in response to hydrostatic pressure.PLoS One. 2014 Sep 12;9(9):e106831. doi: 10.1371/journal.pone.0106831. eCollection 2014. PLoS One. 2014. PMID: 25215865 Free PMC article.
-
P(II) signal transduction proteins are ATPases whose activity is regulated by 2-oxoglutarate.Proc Natl Acad Sci U S A. 2013 Aug 6;110(32):12948-53. doi: 10.1073/pnas.1304386110. Epub 2013 Jul 1. Proc Natl Acad Sci U S A. 2013. PMID: 23818625 Free PMC article.
-
Nutrient-sensing mechanisms across evolution.Cell. 2015 Mar 26;161(1):67-83. doi: 10.1016/j.cell.2015.02.041. Cell. 2015. PMID: 25815986 Free PMC article. Review.
References
-
- Atlante, A., S. Giannattasio, A. Bobba, S. Gagliardi, V. Petragallo, P. Calissano, E. Marra, and S. Passarella. 2005. An increase in the ATP levels occurs in cerebellar granule cells en route to apoptosis in which ATP derives from both oxidative phosphorylation and anaerobic glycolysis. Biochim. Biophys. Acta 170850-62. - PubMed
-
- Bowers, K. C., A. P. Allshire, and P. H. Cobbold. 1992. Bioluminescent measurement in single cardiomyocytes of sudden cytosolic ATP depletion coincident with rigor. J. Mol. Cell Cardiol. 24213-218. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

