Degradome expression profiling in human articular cartilage
- PMID: 19549314
- PMCID: PMC2714152
- DOI: 10.1186/ar2741
Degradome expression profiling in human articular cartilage
Abstract
Introduction: The molecular mechanisms underlying cartilage destruction in osteoarthritis are poorly understood. Proteolysis is a key feature in the turnover and degradation of cartilage extracellular matrix where the focus of research has been on the metzincin family of metalloproteinases. However, there is strong evidence to indicate important roles for other catalytic classes of proteases, with both extracellular and intracellular activities. The aim of this study was to profile the expression of the majority of protease genes in all catalytic classes in normal human cartilage and that from patients with osteoarthritis (OA) using a quantitative method.
Methods: Human cartilage was obtained from femoral heads at joint replacement for either osteoarthritis or following fracture to the neck of femur (NOF). Total RNA was purified, and expression of genes assayed using Taqman low-density array quantitative RT-PCR.
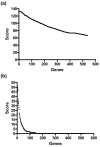
Results: A total of 538 protease genes were profiled, of which 431 were expressed in cartilage. A total of 179 genes were differentially expressed in OA versus NOF cartilage: eight aspartic proteases, 44 cysteine proteases, 76 metalloproteases, 46 serine proteases and five threonine proteases. Wilcoxon ranking as well as the LogitBoost-NR machine learning approach were used to assign significance to each gene, with the most highly ranked genes broadly similar using each method.
Conclusions: This study is the most complete quantitative analysis of protease gene expression in cartilage to date. The data help give direction to future research on the specific function(s) of individual proteases or protease families in cartilage and may help to refine anti-proteolytic strategies in OA.
Figures
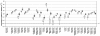
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

