Lutein accumulation in the absence of zeaxanthin restores nonphotochemical quenching in the Arabidopsis thaliana npq1 mutant
- PMID: 19549928
- PMCID: PMC2714924
- DOI: 10.1105/tpc.109.066571
Lutein accumulation in the absence of zeaxanthin restores nonphotochemical quenching in the Arabidopsis thaliana npq1 mutant
Abstract
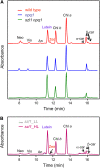
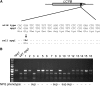
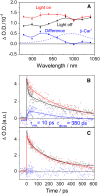
Plants protect themselves from excess absorbed light energy through thermal dissipation, which is measured as nonphotochemical quenching of chlorophyll fluorescence (NPQ). The major component of NPQ, qE, is induced by high transthylakoid DeltapH in excess light and depends on the xanthophyll cycle, in which violaxanthin and antheraxanthin are deepoxidized to form zeaxanthin. To investigate the xanthophyll dependence of qE, we identified suppressor of zeaxanthin-less1 (szl1) as a suppressor of the Arabidopsis thaliana npq1 mutant, which lacks zeaxanthin. szl1 npq1 plants have a partially restored qE but lack zeaxanthin and have low levels of violaxanthin, antheraxanthin, and neoxanthin. However, they accumulate more lutein and alpha-carotene than the wild type. szl1 contains a point mutation in the lycopene beta-cyclase (LCYB) gene. Based on the pigment analysis, LCYB appears to be the major lycopene beta-cyclase and is not involved in neoxanthin synthesis. The Lhcb4 (CP29) and Lhcb5 (CP26) protein levels are reduced by 50% in szl1 npq1 relative to the wild type, whereas other Lhcb proteins are present at wild-type levels. Analysis of carotenoid radical cation formation and leaf absorbance changes strongly suggest that the higher amount of lutein substitutes for zeaxanthin in qE, implying a direct role in qE, as well as a mechanism that is weakly sensitive to carotenoid structural properties.
Figures








Similar articles
-
Arabidopsis mutants define a central role for the xanthophyll cycle in the regulation of photosynthetic energy conversion.Plant Cell. 1998 Jul;10(7):1121-34. doi: 10.1105/tpc.10.7.1121. Plant Cell. 1998. PMID: 9668132 Free PMC article.
-
The zeaxanthin-independent and zeaxanthin-dependent qE components of nonphotochemical quenching involve common conformational changes within the photosystem II antenna in Arabidopsis.Plant Physiol. 2009 Feb;149(2):1061-75. doi: 10.1104/pp.108.129957. Epub 2008 Nov 14. Plant Physiol. 2009. PMID: 19011000 Free PMC article.
-
Zeaxanthin has enhanced antioxidant capacity with respect to all other xanthophylls in Arabidopsis leaves and functions independent of binding to PSII antennae.Plant Physiol. 2007 Dec;145(4):1506-20. doi: 10.1104/pp.107.108480. Epub 2007 Oct 11. Plant Physiol. 2007. PMID: 17932304 Free PMC article.
-
Molecular genetics of xanthophyll-dependent photoprotection in green algae and plants.Philos Trans R Soc Lond B Biol Sci. 2000 Oct 29;355(1402):1385-94. doi: 10.1098/rstb.2000.0700. Philos Trans R Soc Lond B Biol Sci. 2000. PMID: 11127993 Free PMC article. Review.
-
Genetic manipulation of carotenoid biosynthesis and photoprotection.Philos Trans R Soc Lond B Biol Sci. 2000 Oct 29;355(1402):1395-403. doi: 10.1098/rstb.2000.0701. Philos Trans R Soc Lond B Biol Sci. 2000. PMID: 11127994 Free PMC article. Review.
Cited by
-
A kaleidoscope of photosynthetic antenna proteins and their emerging roles.Plant Physiol. 2022 Jun 27;189(3):1204-1219. doi: 10.1093/plphys/kiac175. Plant Physiol. 2022. PMID: 35512089 Free PMC article.
-
Identification of growth regulators using cross-species network analysis in plants.Plant Physiol. 2022 Nov 28;190(4):2350-2365. doi: 10.1093/plphys/kiac374. Plant Physiol. 2022. PMID: 35984294 Free PMC article.
-
Photoprotective Strategies of Mediterranean Plants in Relation to Morphological Traits and Natural Environmental Pressure: A Meta-Analytical Approach.Front Plant Sci. 2017 Jun 19;8:1051. doi: 10.3389/fpls.2017.01051. eCollection 2017. Front Plant Sci. 2017. PMID: 28674548 Free PMC article.
-
Comparison of different cells of Haematococcus pluvialis reveals an extensive acclimation mechanism during its aging process: from a perspective of photosynthesis.PLoS One. 2013 Jul 26;8(7):e67028. doi: 10.1371/journal.pone.0067028. Print 2013. PLoS One. 2013. PMID: 23922648 Free PMC article.
-
Decreased photochemical efficiency of photosystem II following sunlight exposure of shade-grown leaves of avocado: because of, or in spite of, two kinetically distinct xanthophyll cycles?Plant Physiol. 2013 Feb;161(2):836-52. doi: 10.1104/pp.112.209692. Epub 2012 Dec 4. Plant Physiol. 2013. PMID: 23213134 Free PMC article.
References
-
- Ahn, T.K., Avenson, T.J., Ballottari, M., Cheng, Y.C., Niyogi, K.K., Bassi, R., and Fleming, G.R. (2008). Architecture of a charge-transfer state regulating light harvesting in a plant antenna protein. Science 320 794–797. - PubMed
-
- Al-Babili, S., Hugueney, P., Schledz, M., Welsch, R., Frohnmeyer, H., Laule, O., and Beyer, P. (2000). Identification of a novel gene coding for neoxanthin synthase from Solanum tuberosum. FEBS Lett. 485 168–172. - PubMed
-
- Amarie, S., Standfuss, J., Barros, T., Kuhlbrandt, W., Dreuw, A., and Wachtveitl, J. (2007). Carotenoid radical cations as a probe for the molecular mechanism of nonphotochemical quenching in oxygenic photosynthesis. J. Phys. Chem. 111 3481–3487. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

