In silico analysis of the cyclophilin repertoire of apicomplexan parasites
- PMID: 19555495
- PMCID: PMC2713222
- DOI: 10.1186/1756-3305-2-27
In silico analysis of the cyclophilin repertoire of apicomplexan parasites
Abstract
Background: Cyclophilins (Cyps) are peptidyl cis/trans isomerases implicated in diverse processes such as protein folding, signal transduction, and RNA processing. They are also candidate drug targets, in particular for the immunosuppressant cyclosporine A. In addition, cyclosporine is known to exhibit anti-parasitic effects on a wide range of organisms including several apicomplexa. In order to obtain new non-immunosuppressive drugs targeting apicomplexan cyclophilins, a profound knowledge of the cyclophilin repertoire of this phylum would be necessary.
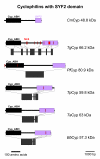
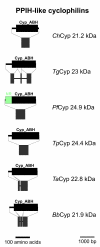
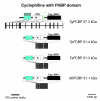
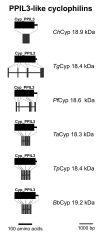
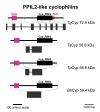
Results: BLAST and maximum likelihood analyses identified 16 different cyclophilin subfamilies within the genomes of Cryptosporidium hominis, Toxoplasma gondii, Plasmodium falciparum, Theileria annulata, Theileria parva, and Babesia bovis. In addition to good statistical support from the phylogenetic analysis, these subfamilies are also confirmed by comparison of cyclophilin domain architecture. Within an individual genome, the number of different Cyp genes that could be deduced varies between 7-9 for Cryptosporidia and 14 for T. gondii. Many of the putative apicomplexan cyclophilins are predicted to be nuclear proteins, most of them presumably involved in RNA processing.
Conclusion: The genomes of apicomplexa harbor a cyclophilin repertoire that is at least as complex as that of most fungi. The identification of Cyp subfamilies that are specific for lower eukaryotes, apicomplexa, or even the genus Plasmodium is of particular interest since these subfamilies are not present in host cells and might therefore represent attractive drug targets.
Figures








Similar articles
-
Divergence of the mitochondrial genome structure in the apicomplexan parasites, Babesia and Theileria.Mol Biol Evol. 2010 May;27(5):1107-16. doi: 10.1093/molbev/msp320. Epub 2009 Dec 24. Mol Biol Evol. 2010. PMID: 20034997
-
A genome-wide analysis of coatomer protein (COP) subunits of apicomplexan parasites and their evolutionary relationships.BMC Genomics. 2019 Jan 31;20(1):98. doi: 10.1186/s12864-019-5463-1. BMC Genomics. 2019. PMID: 30704415 Free PMC article.
-
Babesia gibsoni endemic to Wuhan, China: mitochondrial genome sequencing, annotation, and comparison with apicomplexan parasites.Parasitol Res. 2019 Jan;118(1):235-243. doi: 10.1007/s00436-018-6158-2. Epub 2018 Nov 24. Parasitol Res. 2019. PMID: 30474737
-
Phylogeny and evolution of apicoplasts and apicomplexan parasites.Parasitol Int. 2015 Jun;64(3):254-9. doi: 10.1016/j.parint.2014.10.005. Epub 2014 Oct 14. Parasitol Int. 2015. PMID: 25451217 Review.
-
Global protein expression analysis in apicomplexan parasites: current status.Proteomics. 2005 Mar;5(4):918-24. doi: 10.1002/pmic.200401161. Proteomics. 2005. PMID: 15759314 Review.
Cited by
-
Targeting Artemisinin-Resistant Malaria by Repurposing the Anti-Hepatitis C Virus Drug Alisporivir.Antimicrob Agents Chemother. 2022 Dec 20;66(12):e0039222. doi: 10.1128/aac.00392-22. Epub 2022 Nov 14. Antimicrob Agents Chemother. 2022. PMID: 36374050 Free PMC article.
-
Insights into Peptidyl-Prolyl cis-trans Isomerases from Clinically Important Protozoans: From Structure to Potential Biotechnological Applications.Pathogens. 2024 Jul 31;13(8):644. doi: 10.3390/pathogens13080644. Pathogens. 2024. PMID: 39204244 Free PMC article. Review.
-
The immunophilin repertoire of Plasmodiophora brassicae and functional analysis of PbCYP3 cyclophilin.Mol Genet Genomics. 2018 Apr;293(2):381-390. doi: 10.1007/s00438-017-1395-0. Epub 2017 Nov 11. Mol Genet Genomics. 2018. PMID: 29128880 Free PMC article.
-
Evaluating the potential of non-immunosuppressive cyclosporin analogs for targeting Toxoplasma gondii cyclophilin: Insights from structural studies.Protein Sci. 2024 Oct;33(10):e5157. doi: 10.1002/pro.5157. Protein Sci. 2024. PMID: 39312281 Free PMC article.
-
Cyclophilin A: a key player for etiological agent infection.Appl Microbiol Biotechnol. 2021 Feb;105(4):1365-1377. doi: 10.1007/s00253-021-11115-2. Epub 2021 Jan 25. Appl Microbiol Biotechnol. 2021. PMID: 33492451 Free PMC article. Review.
References
-
- Maruyama T, Furutani M. Archaeal peptidyl prolyl cis-trans isomerases (PPIases) Front Biosci. 2000;5:D821–836. - PubMed
-
- Fischer G, Aumüller T. Regulation of peptide bond cis/trans isomerization by enzyme catalysis and its implication in physiological processes. Reviews of physiology, biochemistry and pharmacology. 2003;148:105–150. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

