Truncation in the tcdC region of the Clostridium difficile PathLoc of clinical isolates does not predict increased biological activity of Toxin B or Toxin A
- PMID: 19558711
- PMCID: PMC2707377
- DOI: 10.1186/1471-2334-9-103
Truncation in the tcdC region of the Clostridium difficile PathLoc of clinical isolates does not predict increased biological activity of Toxin B or Toxin A
Abstract
Background: The increased severity of disease associated with the NAP1 strain of Clostridium difficile has been attributed to mutations to the tcdC gene which codes for a negative regulator of toxin production. To assess the role of hyper-production of Toxins A and B in clinical isolates of Clostridium difficile, two NAP1-related and five NAP1 non-related strains were compared.
Methods: Sequencing was performed on tcdC, tcdR, and tcdE to determine if there were differences that might account for hyper-production of Toxin A and Toxin B in NAP1-related strains. Biological activity of Toxin B was evaluated using the HFF cell CPE assay and Toxin A biological activity was assessed using the Caco-2 Trans-membrane resistance assay.
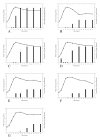
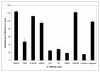
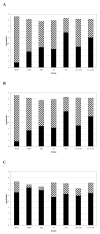
Results: Our results confirm that Toxin A and Toxin B production in NAP1-related strains and ATCC 43255 occurs earlier in the exponential growth phase compared to most NAP1-nonrelated clinical isolates. Despite the hyper-production observed in ATCC 43255 it had no mutations in tcdC, tcdR or tcdE. Analysis of the other clinical isolates indicated that the kinetics and ultimate final concentration of Toxin A and B did not correlate with the presence or lack of alterations in tcdC, tcdR or tcdE.
Conclusion: Our data do not support a direct role for alterations in the tcdC gene as a predictor of hyperproduction of Toxin A and B in NAP1-related strains.
Figures


Similar articles
-
TcdC does not significantly repress toxin expression in Clostridium difficile 630ΔErm.PLoS One. 2012;7(8):e43247. doi: 10.1371/journal.pone.0043247. Epub 2012 Aug 17. PLoS One. 2012. PMID: 22912837 Free PMC article.
-
Human hypervirulent Clostridium difficile strains exhibit increased sporulation as well as robust toxin production.J Bacteriol. 2010 Oct;192(19):4904-11. doi: 10.1128/JB.00445-10. Epub 2010 Jul 30. J Bacteriol. 2010. PMID: 20675495 Free PMC article.
-
Molecular analysis of the pathogenicity locus and polymorphism in the putative negative regulator of toxin production (TcdC) among Clostridium difficile clinical isolates.J Clin Microbiol. 2002 Sep;40(9):3470-5. doi: 10.1128/JCM.40.9.3470-3475.2002. J Clin Microbiol. 2002. PMID: 12202595 Free PMC article.
-
Clostridium difficile toxin synthesis is negatively regulated by TcdC.J Med Microbiol. 2008 Jun;57(Pt 6):685-689. doi: 10.1099/jmm.0.47775-0. J Med Microbiol. 2008. PMID: 18480323 Review.
-
Regulation of Clostridioides difficile toxin production.Curr Opin Microbiol. 2022 Feb;65:95-100. doi: 10.1016/j.mib.2021.10.018. Epub 2021 Nov 12. Curr Opin Microbiol. 2022. PMID: 34781095 Free PMC article. Review.
Cited by
-
TcdC does not significantly repress toxin expression in Clostridium difficile 630ΔErm.PLoS One. 2012;7(8):e43247. doi: 10.1371/journal.pone.0043247. Epub 2012 Aug 17. PLoS One. 2012. PMID: 22912837 Free PMC article.
-
Using phenotype microarrays to determine culture conditions that induce or repress toxin production by Clostridium difficile and other microorganisms.PLoS One. 2013;8(2):e56545. doi: 10.1371/journal.pone.0056545. Epub 2013 Feb 20. PLoS One. 2013. PMID: 23437164 Free PMC article.
-
Emergence of Clinical Clostridioides difficile Isolates With Decreased Susceptibility to Vancomycin.Clin Infect Dis. 2022 Jan 7;74(1):120-126. doi: 10.1093/cid/ciaa912. Clin Infect Dis. 2022. PMID: 35016207 Free PMC article.
-
Fourteen-genome comparison identifies DNA markers for severe-disease-associated strains of Clostridium difficile.J Clin Microbiol. 2011 Jun;49(6):2230-8. doi: 10.1128/JCM.00391-11. Epub 2011 Apr 20. J Clin Microbiol. 2011. PMID: 21508155 Free PMC article.
-
Multilocus Variable-Number Tandem-Repeat Analysis of Clostridioides difficile Clusters in Ribotype 027 Isolates and Lack of Association with Clinical Outcomes.J Clin Microbiol. 2019 Apr 26;57(5):e01724-18. doi: 10.1128/JCM.01724-18. Print 2019 May. J Clin Microbiol. 2019. PMID: 30760531 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

