Structure and uncoating of immature adenovirus
- PMID: 19563809
- PMCID: PMC2749003
- DOI: 10.1016/j.jmb.2009.06.057
Structure and uncoating of immature adenovirus
Abstract
Maturation via proteolytic processing is a common trait in the viral world and is often accompanied by large conformational changes and rearrangements in the capsid. The adenovirus protease has been shown to play a dual role in the viral infectious cycle: (a) in maturation, as viral assembly starts with precursors to several of the structural proteins but ends with proteolytically processed versions in the mature virion, and (b) in entry, because protease-impaired viruses have difficulties in endosome escape and uncoating. Indeed, viruses that have not undergone proteolytic processing are not infectious. We studied the three-dimensional structure of immature adenovirus particles as represented by the adenovirus type 2 thermosensitive mutant ts1 grown under non-permissive conditions and compared it with the mature capsid. Our three-dimensional electron microscopy maps at subnanometer resolution indicate that adenovirus maturation does not involve large-scale conformational changes in the capsid. Difference maps reveal the locations of unprocessed peptides pIIIa and pVI and help define their role in capsid assembly and maturation. An intriguing difference appears in the core, indicating a more compact organization and increased stability of the immature cores. We have further investigated these properties by in vitro disassembly assays. Fluorescence and electron microscopy experiments reveal differences in the stability and uncoating of immature viruses, both at the capsid and core levels, as well as disassembly intermediates not previously imaged.
Figures



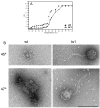
References
-
- Zubieta C, Schoehn G, Chroboczek J, Cusack S. The structure of the human adenovirus 2 penton. Mol Cell. 2005;17:121–35. - PubMed
-
- van Raaij MJ, Mitraki A, Lavigne G, Cusack S. A triple β-spiral in the adenovirus fibre shaft reveals a new structural motif for a fibrous protein. Nature. 1999;401:935–8. - PubMed
-
- Xia D, Henry LJ, Gerard RD, Deisenhofer J. Crystal structure of the receptor-binding domain of adenovirus type 5 fiber protein at 1.7 Å resolution. Structure. 1994;2:1259–70. - PubMed

