Probing the differential interactions of quinazolinedione PD 0305970 and quinolones with gyrase and topoisomerase IV
- PMID: 19564360
- PMCID: PMC2737838
- DOI: 10.1128/AAC.00113-09
Probing the differential interactions of quinazolinedione PD 0305970 and quinolones with gyrase and topoisomerase IV
Abstract
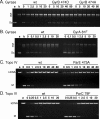
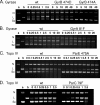
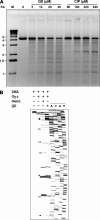
Quinazoline-2,4-diones, such as PD 0305970, are new DNA gyrase and topoisomerase IV (topo IV) inhibitors with potent activity against gram-positive pathogens, including quinolone-resistant isolates. The mechanistic basis of dione activity vis-à-vis quinolones is not understood. We present evidence for Streptococcus pneumoniae gyrase and topo IV that PD 0305970 and quinolones interact differently with the enzyme breakage-reunion and Toprim domains, DNA, and Mg2+-four components that are juxtaposed in the topoisomerase cleavage complex to effect DNA scission. First, PD 0305970 targets primarily gyrase in Streptococcus pneumoniae. However, unlike quinolones, which select predominantly for gyrA (or topo IV parC) mutations in the breakage-reunion domain, unusually the dione selected for novel mutants with alterations that map to a region of the Toprim domain of GyrB (R456H and E474A or E474D) or ParE (D435H and E475A). This "dione resistance-determining region" overlaps the GyrB quinolone resistance-determining region and the region that binds essential Mg2+ ions, each function involving conserved EGDSA and PLRGK motifs. Second, dione-resistant gyrase and topo IV were inhibited by ciprofloxacin, whereas quinolone-resistant enzymes (GyrA S81F and ParC S79F) remained susceptible to PD 0305970. Third, dione-promoted DNA cleavage by gyrase occurred at a distinct repertoire of sites, implying that structural differences with quinolones are sensed at the DNA level. Fourth, unlike the situation with quinolones, the Mg2+ chelator EDTA did not reverse dione-induced gyrase cleavage nor did the dione promote Mg2+-dependent DNA unwinding. It appears that PD 0305970 interacts uniquely to stabilize the cleavage complex of gyrase/topo IV perhaps via an altered orientation directed by the bidentate 3-amino-2,4-dione moiety.
Figures



Similar articles
-
Ciprofloxacin dimers target gyrase in Streptococcus pneumoniae.Antimicrob Agents Chemother. 2004 Jun;48(6):2108-15. doi: 10.1128/AAC.48.6.2108-2115.2004. Antimicrob Agents Chemother. 2004. PMID: 15155208 Free PMC article.
-
Involvement of topoisomerase IV and DNA gyrase as ciprofloxacin targets in Streptococcus pneumoniae.Antimicrob Agents Chemother. 1996 Oct;40(10):2321-6. doi: 10.1128/AAC.40.10.2321. Antimicrob Agents Chemother. 1996. PMID: 8891138 Free PMC article.
-
Cleavable-complex formation by wild-type and quinolone-resistant Streptococcus pneumoniae type II topoisomerases mediated by gemifloxacin and other fluoroquinolones.Antimicrob Agents Chemother. 2002 Feb;46(2):413-9. doi: 10.1128/AAC.46.2.413-419.2002. Antimicrob Agents Chemother. 2002. PMID: 11796351 Free PMC article.
-
Quinolone mode of action.Drugs. 1995;49 Suppl 2:10-5. doi: 10.2165/00003495-199500492-00004. Drugs. 1995. PMID: 8549276 Review.
-
DNA Topoisomerase Inhibitors: Trapping a DNA-Cleaving Machine in Motion.J Mol Biol. 2019 Aug 23;431(18):3427-3449. doi: 10.1016/j.jmb.2019.07.008. Epub 2019 Jul 10. J Mol Biol. 2019. PMID: 31301408 Free PMC article. Review.
Cited by
-
Impact of amino acid substitutions in B subunit of DNA gyrase in Mycobacterium leprae on fluoroquinolone resistance.PLoS Negl Trop Dis. 2012;6(10):e1838. doi: 10.1371/journal.pntd.0001838. Epub 2012 Oct 11. PLoS Negl Trop Dis. 2012. PMID: 23071850 Free PMC article.
-
Trapping of the transport-segment DNA by the ATPase domains of a type II topoisomerase.Nat Commun. 2018 Jul 3;9(1):2579. doi: 10.1038/s41467-018-05005-x. Nat Commun. 2018. PMID: 29968711 Free PMC article.
-
Role of the water-metal ion bridge in mediating interactions between quinolones and Escherichia coli topoisomerase IV.Biochemistry. 2014 Sep 2;53(34):5558-67. doi: 10.1021/bi500682e. Epub 2014 Aug 21. Biochemistry. 2014. PMID: 25115926 Free PMC article.
-
Crystal structure and stability of gyrase-fluoroquinolone cleaved complexes from Mycobacterium tuberculosis.Proc Natl Acad Sci U S A. 2016 Feb 16;113(7):1706-13. doi: 10.1073/pnas.1525047113. Epub 2016 Jan 20. Proc Natl Acad Sci U S A. 2016. PMID: 26792525 Free PMC article.
-
Ubiquitous Nature of Fluoroquinolones: The Oscillation between Antibacterial and Anticancer Activities.Antibiotics (Basel). 2017 Nov 7;6(4):26. doi: 10.3390/antibiotics6040026. Antibiotics (Basel). 2017. PMID: 29112154 Free PMC article. Review.
References
-
- Alovero, F. L., X.-S. Pan, J. E. Morris, R. H. Manzo, and L. M. Fisher. 2000. Engineering the specificity of antibacterial fluoroquinolones: benzenesulfonamide modifications at C-7 of ciprofloxacin change its primary target in Streptococcus pneumoniae from topoisomerase IV to gyrase. Antimicrob. Agents Chemother. 44:320-325. - PMC - PubMed
-
- Bassetti, M., G. Medica, A. Di Biagio, E. Right, R. Rosso, and D. Bassetti. 2004. New antibiotics for treatment of serious infections due to resistant gram-positive cocci. Rev. Med. Microbiol. 15:109-117.
-
- Bellon, S., J. D. Parsons, Y. Wei, K. Hayakawa, L. L. Swenson, P. S. Charifson, J. A. Lippke, R. Aldape, and C. H. Gross. 2004. Crystal structures of Escherichia coli topoisomerase IV ParE subunit (24 and 43 kilodaltons): a single residue dictates differences in novobiocin potency against topoisomerase IV and DNA gyrase. Antimicrob. Agents Chemother. 48:1856-1864. - PMC - PubMed
-
- Berger, J. M., S. J. Gamblin, S. C. Harrison, and J. C. Wang. 1996. Structure and mechanism of DNA topoisomerase II. Nature 379:225-232. - PubMed
-
- Champoux, J. J. 2001. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70:369-413. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

