Differential regulation of elastic fiber formation by fibulin-4 and -5
- PMID: 19570982
- PMCID: PMC2782046
- DOI: 10.1074/jbc.M109.019364
Differential regulation of elastic fiber formation by fibulin-4 and -5
Abstract
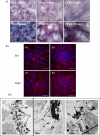
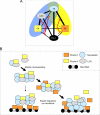
Fibulin-4 and -5 are extracellular glycoproteins with essential non-compensatory roles in elastic fiber assembly. We have determined how they interact with tropoelastin, lysyl oxidase, and fibrillin-1, thereby revealing how they differentially regulate assembly. Strong binding between fibulin-4 and lysyl oxidase enhanced the interaction of fibulin-4 with tropoelastin, forming ternary complexes that may direct elastin cross-linking. In contrast, fibulin-5 did not bind lysyl oxidase strongly but bound tropoelastin in terminal and central regions and could concurrently bind fibulin-4. Both fibulins differentially bound N-terminal fibrillin-1, which strongly inhibited their binding to lysyl oxidase and tropoelastin. Knockdown experiments revealed that fibulin-5 controlled elastin deposition on microfibrils, although fibulin-4 can also bind fibrillin-1. These experiments provide a molecular account of the distinct roles of fibulin-4 and -5 in elastic fiber assembly and how they act in concert to chaperone cross-linked elastin onto microfibrils.
Figures



Similar articles
-
DANCE/fibulin-5 promotes elastic fiber formation in a tropoelastin isoform-dependent manner.Clin Biochem. 2009 May;42(7-8):713-21. doi: 10.1016/j.clinbiochem.2008.12.020. Epub 2009 Jan 8. Clin Biochem. 2009. PMID: 19167375
-
LTBP-2 competes with tropoelastin for binding to fibulin-5 and heparin, and is a negative modulator of elastinogenesis.Matrix Biol. 2014 Feb;34:114-23. doi: 10.1016/j.matbio.2013.10.007. Epub 2013 Oct 19. Matrix Biol. 2014. PMID: 24148803
-
Characterization of Microfibrillar-associated Protein 4 (MFAP4) as a Tropoelastin- and Fibrillin-binding Protein Involved in Elastic Fiber Formation.J Biol Chem. 2016 Jan 15;291(3):1103-14. doi: 10.1074/jbc.M115.681775. Epub 2015 Nov 24. J Biol Chem. 2016. PMID: 26601954 Free PMC article.
-
New insights into elastic fiber assembly.Birth Defects Res C Embryo Today. 2007 Dec;81(4):229-40. doi: 10.1002/bdrc.20111. Birth Defects Res C Embryo Today. 2007. PMID: 18228265 Review.
-
Elastic tissue disruption is a major pathogenic factor to human vascular disease.Mol Biol Rep. 2021 May;48(5):4865-4878. doi: 10.1007/s11033-021-06478-8. Epub 2021 Jun 15. Mol Biol Rep. 2021. PMID: 34129188 Review.
Cited by
-
Stem cell derived extracellular vesicles for vascular elastic matrix regenerative repair.Acta Biomater. 2020 Sep 1;113:267-278. doi: 10.1016/j.actbio.2020.07.002. Epub 2020 Jul 6. Acta Biomater. 2020. PMID: 32645438 Free PMC article.
-
Alternative splicing and tissue-specific elastin misassembly act as biological modifiers of human elastin gene frameshift mutations associated with dominant cutis laxa.J Biol Chem. 2012 Jun 22;287(26):22055-67. doi: 10.1074/jbc.M111.327940. Epub 2012 May 9. J Biol Chem. 2012. PMID: 22573328 Free PMC article.
-
EFEMP2 Inhibits Breast Cancer Invasion And Metastasis In Vitro And In Vivo.Onco Targets Ther. 2019 Oct 30;12:8915-8933. doi: 10.2147/OTT.S221219. eCollection 2019. Onco Targets Ther. 2019. PMID: 31802903 Free PMC article.
-
Inhibitory effects of cycloastragenol on abdominal aortic aneurysm and its related mechanisms.Br J Pharmacol. 2019 Jan;176(2):282-296. doi: 10.1111/bph.14515. Epub 2018 Dec 4. Br J Pharmacol. 2019. PMID: 30302749 Free PMC article.
-
The "Elastic Perspective" of SARS-CoV-2 Infection and the Role of Intrinsic and Extrinsic Factors.Int J Mol Sci. 2022 Jan 29;23(3):1559. doi: 10.3390/ijms23031559. Int J Mol Sci. 2022. PMID: 35163482 Free PMC article. Review.
References
-
- Chu M. L., Tsuda T. (2004) Birth Defects Res. C Embryo Today 72, 25–36 - PubMed
-
- Timpl R., Sasaki T., Kostka G., Chu M. L. (2003) Nat. Rev. Mol. Cell Biol. 4, 479–489 - PubMed
-
- Kobayashi N., Kostka G., Garbe J. H., Keene D. R., Bächinger H. P., Hanisch F. G., Markova D., Tsuda T., Timpl R., Chu M. L., Sasaki T. (2007) J. Biol. Chem. 282, 11805–11816 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

