Phylogenetic analysis identifies many uncharacterized actin-like proteins (Alps) in bacteria: regulated polymerization, dynamic instability and treadmilling in Alp7A
- PMID: 19602153
- PMCID: PMC2814180
- DOI: 10.1111/j.1365-2958.2009.06771.x
Phylogenetic analysis identifies many uncharacterized actin-like proteins (Alps) in bacteria: regulated polymerization, dynamic instability and treadmilling in Alp7A
Abstract
Actin, one of the most abundant proteins in the eukaryotic cell, also has an abundance of relatives in the eukaryotic proteome. To date though, only five families of actins have been characterized in bacteria. We have conducted a phylogenetic search and uncovered more than 35 highly divergent families of actin-like proteins (Alps) in bacteria. Their genes are found primarily on phage genomes, on plasmids and on integrating conjugative elements, and are likely to be involved in a variety of functions. We characterize three Alps and find that all form filaments in the cell. The filaments of Alp7A, a plasmid partitioning protein and one of the most divergent of the Alps, display dynamic instability and also treadmill. Alp7A requires other elements from the plasmid to assemble into dynamic polymers in the cell. Our findings suggest that most if not all of the Alps are indeed actin relatives, and that actin is very well represented in bacteria.
Figures





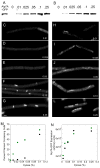

References
-
- Belmont LD, Patterson GM, Drubin DG. New actin mutants allow further characterization of the nucleotide binding cleft and drug binding sites. J Cell Sci. 1999;112:1325–1336. - PubMed
-
- Bingham JB, Schroer TA. Self-regulated polymerization of the actin-related protein Arp1. Curr Biol. 1999;9:223–226. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

