Mammalian polyamine metabolism and function
- PMID: 19603518
- PMCID: PMC2753421
- DOI: 10.1002/iub.230
Mammalian polyamine metabolism and function
Abstract
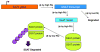
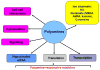
Polyamines are ubiquitous small basic molecules that play multiple essential roles in mammalian physiology. Their cellular content is highly regulated and there is convincing evidence that altered metabolism is involvement in many disease states. Drugs altering polyamine levels may therefore have a variety of important targets. This review will summarize the current state of understanding of polyamine metabolism and function, the regulation of polyamine content, and heritable pathological conditions that may be derived from altered polyamine metabolism.
(c) 2009 IUBMB
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

