Quantitative epigenetics: DNA sequence variation need not apply
- PMID: 19605680
- PMCID: PMC3959989
- DOI: 10.1101/gad.1824909
Quantitative epigenetics: DNA sequence variation need not apply
Abstract
Two recent reports, including one by Reinders and colleagues (pp. 939-950) in the April 15, 2009, issue of Genes & Development, describe the construction of Arabidopsis recombinant inbred populations that maximize epigenetic rather than genetic variation. The distribution and behavior of phenotypic variation in these populations suggest that stable epialleles can control complex quantitative traits. However, stochastic epimutation and transposon movement in these populations present some unexpected technical hurdles to implementing quantitative epigenetic analysis.
Figures
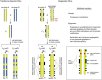
References
-
- Bossdorf O, Richards CL, Pigliucci M. Epigenetics for ecologists. Ecol Lett. 2008;11:106–115. - PubMed
-
- Chen RZ, Pettersson U, Beard C, Jackson-Grusby L, Jaenisch R. DNA hypomethylation leads to elevated mutation rates. Nature. 1998;395:89–93. - PubMed
-
- Cubas P, Vincent C, Coen E. An epigenetic mutation responsible for natural variation in floral symmetry. Nature. 1999;401:157–161. - PubMed
-
- Gal-Yam EN, Saito Y, Egger G, Jones PA. Cancer epigenetics: Modifications, screening, and therapy. Annu Rev Med. 2008;59:267–280. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
