Global stabilization of rRNA structure by ribosomal proteins S4, S17, and S20
- PMID: 19616559
- PMCID: PMC2763185
- DOI: 10.1016/j.jmb.2009.07.032
Global stabilization of rRNA structure by ribosomal proteins S4, S17, and S20
Abstract
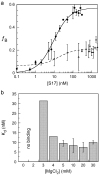
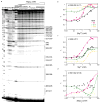
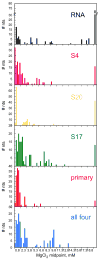
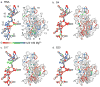
Ribosomal proteins stabilize the folded structure of the ribosomal RNA and enable the recruitment of further proteins to the complex. Quantitative hydroxyl radical footprinting was used to measure the extent to which three different primary assembly proteins, S4, S17, and S20, stabilize the three-dimensional structure of the Escherichia coli 16S 5' domain. The stability of the complexes was perturbed by varying the concentration of MgCl(2). Each protein influences the stability of the ribosomal RNA tertiary interactions beyond its immediate binding site. S4 and S17 stabilize the entire 5' domain, while S20 has a more local effect. Multistage folding of individual helices within the 5' domain shows that each protein stabilizes a different ensemble of structural intermediates that include nonnative interactions at low Mg(2+) concentration. We propose that the combined interactions of S4, S17, and S20 with different helical junctions bias the free-energy landscape toward a few RNA conformations that are competent to add the secondary assembly protein S16 in the next step of assembly.
Figures
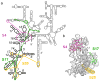
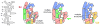
Similar articles
-
Differential effects of ribosomal proteins and Mg2+ ions on a conformational switch during 30S ribosome 5'-domain assembly.RNA. 2015 Nov;21(11):1859-65. doi: 10.1261/rna.051292.115. Epub 2015 Sep 9. RNA. 2015. PMID: 26354770 Free PMC article.
-
S16 throws a conformational switch during assembly of 30S 5' domain.Nat Struct Mol Biol. 2009 Apr;16(4):438-45. doi: 10.1038/nsmb.1585. Epub 2009 Apr 3. Nat Struct Mol Biol. 2009. PMID: 19343072 Free PMC article.
-
Interaction of proteins S16, S17 and S20 with 16 S ribosomal RNA.J Mol Biol. 1988 Mar 20;200(2):291-9. doi: 10.1016/0022-2836(88)90241-0. J Mol Biol. 1988. PMID: 3373529
-
Assembly of bacterial ribosomes.Annu Rev Biochem. 2011;80:501-26. doi: 10.1146/annurev-biochem-062608-160432. Annu Rev Biochem. 2011. PMID: 21529161 Review.
-
Structure and dynamics of ribosomal RNA.Curr Opin Struct Biol. 1998 Jun;8(3):294-300. doi: 10.1016/s0959-440x(98)80061-4. Curr Opin Struct Biol. 1998. PMID: 9666324 Review.
Cited by
-
The long-range P3 helix of the Tetrahymena ribozyme is disrupted during folding between the native and misfolded conformations.J Mol Biol. 2013 Aug 9;425(15):2670-86. doi: 10.1016/j.jmb.2013.05.008. Epub 2013 May 20. J Mol Biol. 2013. PMID: 23702292 Free PMC article.
-
Protein-guided RNA dynamics during early ribosome assembly.Nature. 2014 Feb 20;506(7488):334-8. doi: 10.1038/nature13039. Epub 2014 Feb 12. Nature. 2014. PMID: 24522531 Free PMC article.
-
Comparative Analysis of Structural and Dynamical Features of Ribosome Upon Association With mRNA Reveals Potential Role of Ribosomal Proteins.Front Mol Biosci. 2021 Aug 2;8:654164. doi: 10.3389/fmolb.2021.654164. eCollection 2021. Front Mol Biosci. 2021. PMID: 34409066 Free PMC article.
-
Structurally heterogeneous ribosomes cooperate in protein synthesis in bacterial cells.Nat Commun. 2025 Mar 20;16(1):2751. doi: 10.1038/s41467-025-57955-8. Nat Commun. 2025. PMID: 40113756 Free PMC article.
-
Differential effects of ribosomal proteins and Mg2+ ions on a conformational switch during 30S ribosome 5'-domain assembly.RNA. 2015 Nov;21(11):1859-65. doi: 10.1261/rna.051292.115. Epub 2015 Sep 9. RNA. 2015. PMID: 26354770 Free PMC article.
References
-
- Nierhaus KH. The assembly of prokaryotic ribosomes. Biochimie. 1991;73:739–755. - PubMed
-
- Warner JR, Vilardell J, Sohn JH. Economics of ribosome biosynthesis. Cold Spring Harb Symp Quant Biol. 2001;66:567–574. - PubMed
-
- Mizushima S, Nomura M. Assembly mapping of 30S ribosomal proteins from E. coli. Nature. 1970;226:1214. - PubMed
-
- Stern S, Powers T, Changchien LM, Noller HF. RNA-protein interactions in 30S ribosomal subunits: folding and function of 16S rRNA. Science. 1989;244:783–790. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

