The Drosophila U7 snRNP proteins Lsm10 and Lsm11 are required for histone pre-mRNA processing and play an essential role in development
- PMID: 19620235
- PMCID: PMC2743060
- DOI: 10.1261/rna.1518009
The Drosophila U7 snRNP proteins Lsm10 and Lsm11 are required for histone pre-mRNA processing and play an essential role in development
Abstract
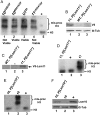
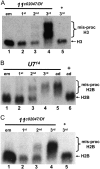
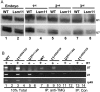
Metazoan replication-dependent histone mRNAs are not polyadenylated, and instead terminate in a conserved stem-loop structure generated by an endonucleolytic cleavage of the pre-mRNA involving U7 snRNP. U7 snRNP contains two like-Sm proteins, Lsm10 and Lsm11, which replace SmD1 and SmD2 in the canonical heptameric Sm protein ring that binds spliceosomal snRNAs. Here we show that mutations in either the Drosophila Lsm10 or the Lsm11 gene disrupt normal histone pre-mRNA processing, resulting in production of poly(A)+ histone mRNA as a result of transcriptional read-through to cryptic polyadenylation sites present downstream from each histone gene. This molecular phenotype is indistinguishable from that which we previously described for mutations in U7 snRNA. Lsm10 protein fails to accumulate in Lsm11 mutants, suggesting that a pool of Lsm10-Lsm11 dimers provides precursors for U7 snRNP assembly. Unexpectedly, U7 snRNA was detected in Lsm11 and Lsm1 mutants and could be precipitated with anti-trimethylguanosine antibodies, suggesting that it assembles into a snRNP particle in the absence of Lsm10 and Lsm11. However, this U7 snRNA could not be detected at the histone locus body, suggesting that Lsm10 and Lsm11 are necessary for U7 snRNP localization. In contrast to U7 snRNA null mutants, which are viable, Lsm10 and Lsm11 mutants do not survive to adulthood. Because we cannot detect differences in the histone mRNA phenotype between Lsm10 or Lsm11 and U7 mutants, we propose that the different terminal developmental phenotypes result from the participation of Lsm10 and Lsm11 in an essential function that is distinct from histone pre-mRNA processing and that is independent of U7 snRNA.
Figures




Similar articles
-
Interaction between FLASH and Lsm11 is essential for histone pre-mRNA processing in vivo in Drosophila.RNA. 2011 Jun;17(6):1132-47. doi: 10.1261/rna.2566811. Epub 2011 Apr 27. RNA. 2011. PMID: 21525146 Free PMC article.
-
U7 snRNA mutations in Drosophila block histone pre-mRNA processing and disrupt oogenesis.RNA. 2006 Mar;12(3):396-409. doi: 10.1261/rna.2270406. RNA. 2006. PMID: 16495235 Free PMC article.
-
U7 snRNP is recruited to histone pre-mRNA in a FLASH-dependent manner by two separate regions of the stem-loop binding protein.RNA. 2017 Jun;23(6):938-951. doi: 10.1261/rna.060806.117. Epub 2017 Mar 13. RNA. 2017. PMID: 28289156 Free PMC article.
-
The special Sm core structure of the U7 snRNP: far-reaching significance of a small nuclear ribonucleoprotein.Cell Mol Life Sci. 2004 Oct;61(19-20):2560-70. doi: 10.1007/s00018-004-4190-0. Cell Mol Life Sci. 2004. PMID: 15526162 Free PMC article. Review.
-
U7 deciphered: the mechanism that forms the unusual 3' end of metazoan replication-dependent histone mRNAs.Biochem Soc Trans. 2021 Nov 1;49(5):2229-2240. doi: 10.1042/BST20210323. Biochem Soc Trans. 2021. PMID: 34351387 Free PMC article. Review.
Cited by
-
Drosophila Symplekin localizes dynamically to the histone locus body and tricellular junctions.Nucleus. 2014;5(6):613-25. doi: 10.4161/19491034.2014.990860. Nucleus. 2014. PMID: 25493544 Free PMC article.
-
Interaction between FLASH and Lsm11 is essential for histone pre-mRNA processing in vivo in Drosophila.RNA. 2011 Jun;17(6):1132-47. doi: 10.1261/rna.2566811. Epub 2011 Apr 27. RNA. 2011. PMID: 21525146 Free PMC article.
-
An in vivo genetic screen for genes involved in spliced leader trans-splicing indicates a crucial role for continuous de novo spliced leader RNP assembly.Nucleic Acids Res. 2017 Aug 21;45(14):8474-8483. doi: 10.1093/nar/gkx500. Nucleic Acids Res. 2017. PMID: 28582530 Free PMC article.
-
Drosophila histone locus body assembly and function involves multiple interactions.Mol Biol Cell. 2020 Jul 1;31(14):1525-1537. doi: 10.1091/mbc.E20-03-0176. Epub 2020 May 13. Mol Biol Cell. 2020. PMID: 32401666 Free PMC article.
-
Birth and Death of Histone mRNAs.Trends Genet. 2017 Oct;33(10):745-759. doi: 10.1016/j.tig.2017.07.014. Epub 2017 Aug 31. Trends Genet. 2017. PMID: 28867047 Free PMC article. Review.
References
-
- Beggs JD. Lsm proteins and RNA processing. Biochem Soc Trans. 2005;33:433–438. - PubMed
-
- Bond UM, Yario TA, Steitz JA. Multiple processing-defective mutations in a mammalian histone pre-mRNA are suppressed by compensatory changes in U7 RNA both in vivo and in vitro. Genes & Dev. 1991;5:1709–1722. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
