The N-end rule pathway controls multiple functions during Arabidopsis shoot and leaf development
- PMID: 19620738
- PMCID: PMC2726413
- DOI: 10.1073/pnas.0906404106
The N-end rule pathway controls multiple functions during Arabidopsis shoot and leaf development
Abstract
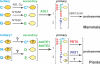
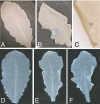
The ubiquitin-dependent N-end rule pathway relates the in vivo half-life of a protein to the identity of its N-terminal residue. This proteolytic system is present in all organisms examined and has been shown to have a multitude of functions in animals and fungi. In plants, however, the functional understanding of the N-end rule pathway is only beginning. The N-end rule has a hierarchic structure. Destabilizing activity of N-terminal Asp, Glu, and (oxidized) Cys requires their conjugation to Arg by an arginyl-tRNA-protein transferase (R-transferase). The resulting N-terminal Arg is recognized by the pathway's E3 ubiquitin ligases, called "N-recognins." Here, we show that the Arabidopsis R-transferases AtATE1 and AtATE2 regulate various aspects of leaf and shoot development. We also show that the previously identified N-recognin PROTEOLYSIS6 (PRT6) mediates these R-transferase-dependent activities. We further demonstrate that the arginylation branch of the N-end rule pathway plays a role in repressing the meristem-promoting BREVIPEDICELLUS (BP) gene in developing leaves. BP expression is known to be excluded from Arabidopsis leaves by the activities of the ASYMMETRIC LEAVES1 (AS1) transcription factor complex and the phytohormone auxin. Our results suggest that AtATE1 and AtATE2 act redundantly with AS1, but independently of auxin, in the control of leaf development.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Bachmair A, Varshavsky A. The degradation signal in a short-lived protein. Cell. 1989;56:1019–1032. - PubMed
-
- Bachmair A, Finley D, Varshavsky A. In vivo half-life of a protein is a function of its amino-terminal residue. Science. 1986;234:179–186. - PubMed
-
- Hu RG, et al. The N-end rule pathway as a nitric oxide sensor controlling the levels of multiple regulators. Nature. 2005;437:981–986. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

