Targeting of the GTPase Irgm1 to the phagosomal membrane via PtdIns(3,4)P(2) and PtdIns(3,4,5)P(3) promotes immunity to mycobacteria
- PMID: 19620982
- PMCID: PMC2715447
- DOI: 10.1038/ni.1759
Targeting of the GTPase Irgm1 to the phagosomal membrane via PtdIns(3,4)P(2) and PtdIns(3,4,5)P(3) promotes immunity to mycobacteria
Abstract
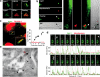
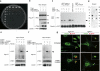
Vertebrate immunity to infection enlists a newly identified family of 47-kilodalton immunity-related GTPases (IRGs). One IRG in particular, Irgm1, is essential for macrophage host defense against phagosomal pathogens, including Mycobacterium tuberculosis (Mtb). Here we show that Irgm1 targets the mycobacterial phagosome through lipid-mediated interactions with phosphatidylinositol-3,4-bisphosphate (PtdIns(3,4)P(2)) and PtdIns(3,4,5)P(3). An isolated Irgm1 amphipathic helix conferred lipid binding in vitro and in vivo. Substitutions in this region blocked phagosome recruitment and failed to complement the antimicrobial defect in Irgm1(-/-) macrophages. Removal of PtdIns(3,4,5)P(3) or inhibition of class I phosphatidylinositol-3-OH kinase (PI(3)K) mimicked this effect in wild-type cells. Cooperation between Irgm1 and PI(3)K further facilitated the engagement of Irgm1 with its fusogenic effectors at the site of infection, thereby ensuring pathogen-directed responses during innate immunity.
Figures




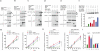
Comment in
-
New insight into the everlasting host-pathogen arms race.Nat Immunol. 2009 Aug;10(8):808-9. doi: 10.1038/ni0809-808. Nat Immunol. 2009. PMID: 19621040 No abstract available.
References
-
- Hingley-Wilson SM, Sambandamurthy VK, Jacobs WR., Jr. Survival perspectives from the world's most successful pathogen, Mycobacterium tuberculosis. Nat. Immunol. 2003;4:949–955. - PubMed
-
- Pieters J. Mycobacterium tuberculosis and the macrophage. Maintaining a balance. Cell Host & Microbe. 2008;3:399–407. - PubMed
-
- MacMicking JD, Taylor GA, McKinney JD. Immune control of tuberculosis by IFN-γ-inducible LRG-47. Science. 2003;302:654–659. - PubMed
-
- Gutierrez MG, et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell. 2004;119:753–766. - PubMed
-
- Singh SB, Davis AS, Taylor GA, Deretic V. Human IRGM induces autophagy to eliminate intracellular bacteria. Science. 2006;313:1438–1441. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

