Gamma-phage lysin PlyG sequence-based synthetic peptides coupled with Qdot-nanocrystals are useful for developing detection methods for Bacillus anthracis by using its surrogates, B. anthracis-Sterne and B. cereus-4342
- PMID: 19624851
- PMCID: PMC2722591
- DOI: 10.1186/1472-6750-9-67
Gamma-phage lysin PlyG sequence-based synthetic peptides coupled with Qdot-nanocrystals are useful for developing detection methods for Bacillus anthracis by using its surrogates, B. anthracis-Sterne and B. cereus-4342
Abstract
Background: Previous reports of site-directed deletion analysis on gamma (gamma)-phage lysin protein (PlyG) have demonstrated that removal of a short amino acid sequence in the C-terminal region encompassing a 10-amino acid motif (190LKMTADFILQ199) abrogates its binding activity specific to the cell wall of Bacillus anthracis. Whether short synthetic peptides representing the10-amino acid PlyG putative binding motif flanked by surrounding N- and C-terminal residues also selectively bind to the bacterial cell wall has not been evaluated. If such peptides do demonstrate selective binding to the cell wall, they could serve as bio-probes towards developing detection technologies for B. anthracis. Furthermore, by using B. anthracis (Sterne, 34F2), an animal vaccine and B. cereus-4342, a gamma-phage susceptible rare strain as surrogates of B. anthracis, development of proof-of-concepts for B. anthracis are feasible.
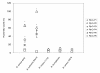
Results: Using four different methods, we evaluated six synthetic peptides representing the putative binding motif including flanking sequences (PlyG-P1 through P6) for the bacterial cell wall binding capacity. Our analysis identified PlyG-P1, PlyG-P3 and PlyG-P5 to have binding capability to both B. anthracis (Sterne, 34F2) and B. cereus-4342. The peptides however did not bind to B. cereus-11778, B. thuringiensis, and B. cereus-10876 suggesting their specificity for B. anthracis-Sterne and B. cereus-4342. PlyG-P3 in combination with fluorescent light microscopy detected even a single bacterium in plasma spiked with the bacteria.
Conclusion: Overall, these studies illustrate that the short 10-amino acid sequence 'LKMTADFILQ' in fact is a stand-alone bacterial cell wall-binding motif of PlyG. In principle, synthetic peptides PlyG-P1, PlyG-P3 and PlyG-P5, especially PlyG-P3 coupled with Qdot-nanocrystals are useful as high-sensitivity bio-probes in developing detection technologies for B. anthracis.
Figures
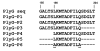


Similar articles
-
Characterization of the catalytic activity of the gamma-phage lysin, PlyG, specific for Bacillus anthracis.FEMS Microbiol Lett. 2008 Sep;286(2):236-40. doi: 10.1111/j.1574-6968.2008.01280.x. Epub 2008 Jul 25. FEMS Microbiol Lett. 2008. PMID: 18662316
-
The secondary cell wall polysaccharide of Bacillus anthracis provides the specific binding ligand for the C-terminal cell wall-binding domain of two phage endolysins, PlyL and PlyG.Glycobiology. 2013 Jul;23(7):820-32. doi: 10.1093/glycob/cwt019. Epub 2013 Mar 14. Glycobiology. 2013. PMID: 23493680 Free PMC article.
-
Peptides panned from a phage-displayed random peptide library are useful for the detection of Bacillus anthracis surrogates B. cereus 4342 and B. anthracis Sterne.Biochem Biophys Res Commun. 2010 Apr 23;395(1):93-8. doi: 10.1016/j.bbrc.2010.03.145. Epub 2010 Mar 27. Biochem Biophys Res Commun. 2010. PMID: 20350526
-
Discovery of phage display peptide ligands for species-specific detection of Bacillus spores.J Microbiol Methods. 2003 May;53(2):263-71. doi: 10.1016/s0167-7012(03)00030-7. J Microbiol Methods. 2003. PMID: 12654497 Review.
-
Characterization and comparative genomic analysis of bacteriophages infecting members of the Bacillus cereus group.Arch Virol. 2014 May;159(5):871-84. doi: 10.1007/s00705-013-1920-3. Epub 2013 Nov 22. Arch Virol. 2014. PMID: 24264384 Review.
Cited by
-
Structure-based modification of a Clostridium difficile-targeting endolysin affects activity and host range.J Bacteriol. 2011 Oct;193(19):5477-86. doi: 10.1128/JB.00439-11. Epub 2011 Jul 29. J Bacteriol. 2011. PMID: 21803993 Free PMC article.
-
Bacteriophage endolysins as novel antimicrobials.Future Microbiol. 2012 Oct;7(10):1147-71. doi: 10.2217/fmb.12.97. Future Microbiol. 2012. PMID: 23030422 Free PMC article. Review.
-
Genomic sequence of bacteriophage ATCC 8074-B1 and activity of its endolysin and engineered variants against Clostridium sporogenes.Appl Environ Microbiol. 2012 May;78(10):3685-92. doi: 10.1128/AEM.07884-11. Epub 2012 Mar 16. Appl Environ Microbiol. 2012. PMID: 22427494 Free PMC article.
-
Endolysin, a Promising Solution against Antimicrobial Resistance.Antibiotics (Basel). 2021 Oct 20;10(11):1277. doi: 10.3390/antibiotics10111277. Antibiotics (Basel). 2021. PMID: 34827215 Free PMC article. Review.
-
Bacillus anthracis gamma phage lysis among soil bacteria: an update on test specificity.BMC Res Notes. 2017 Nov 16;10(1):598. doi: 10.1186/s13104-017-2919-8. BMC Res Notes. 2017. PMID: 29145870 Free PMC article.
References
-
- Borio L, Frank D, Mani V, Chiriboga C, Pollanen M, Ripple M, Ali S, DiAngelo C, Lee J, Arden J, Titus J, Fowler D, O' Toole T, Masur H, Bartlett J, Inglesby T. Death due to bioterrorism-related inhalational anthrax: report of 2 patients. J Am Med Assoc. 2001;286:2554–2559. doi: 10.1001/jama.286.20.2554. - DOI - PubMed
-
- Turnbull PCB. Guidelines for the surveillance and control of anthrax in humans and animals. Third. World Health Organisation. Geneva; 1998. pp. 30–35.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

