Role of cleavage by separase of the Rec8 kleisin subunit of cohesin during mammalian meiosis I
- PMID: 19625504
- PMCID: PMC2909317
- DOI: 10.1242/jcs.035287
Role of cleavage by separase of the Rec8 kleisin subunit of cohesin during mammalian meiosis I
Abstract
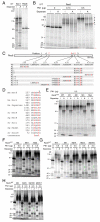
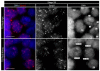
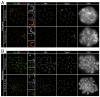
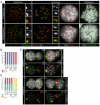
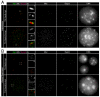
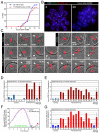
Proteolytic activity of separase is required for chiasma resolution during meiosis I in mouse oocytes. Rec8, the meiosis-specific alpha-kleisin subunit of cohesin, is a key target of separase in yeast. Is the equivalent protein also a target in mammals? We show here that separase cleaves mouse Rec8 at three positions in vitro but only when the latter is hyper-phosphorylated. Expression of a Rec8 variant (Rec8-N) that cannot be cleaved in vitro at these sites causes sterility in male mice. Their seminiferous tubules lack a normal complement of 2 C secondary spermatocytes and 1 C spermatids and contain instead a high proportion of cells with enlarged nuclei. Chromosome spreads reveal that Rec8-N expression has no effect in primary spermatocytes but produces secondary spermatocytes and spermatids with a 4 C DNA content, suggesting that the first and possibly also the second meiotic division is abolished. Expression of Rec8-N in oocytes causes chromosome segregation to be asynchronous and delays its completion by 2-3 hours during anaphase I, probably due to inefficient proteolysis of Rec8-N by separase. Despite this effect, chromosome segregation must be quite accurate as Rec8-N does not greatly reduce female fertility. Our data is consistent with the notion that Rec8 cleavage is important and probably crucial for the resolution of chiasmata in males and females.
Figures



References
-
- Alexandru, G., Uhlmann, F., Mechtler, K., Poupart, M. A. and Nasmyth, K. (2001). Phosphorylation of the cohesin subunit Scc1 by Polo/Cdc5 kinase regulates sister chromatid separation in yeast. Cell 105, 459-472. - PubMed
-
- Bannister, L. A., Reinholdt, L. G., Munroe, R. J. and Schimenti, J. C. (2004). Positional cloning and characterization of mouse mei8, a disrupted allelle of the meiotic cohesin Rec8. Genesis 40, 184-194. - PubMed
-
- Brar, G. A., Kiburz, B. M., Zhang, Y., Kim, J. E., White, F. and Amon, A. (2006). Rec8 phosphorylation and recombination promote the step-wise loss of cohesins in meiosis. Nature 441, 532-536. - PubMed
-
- Buonomo, S. B., Clyne, R. K., Fuchs, J., Loidl, J., Uhlmann, F. and Nasmyth, K. (2000). Disjunction of homologous chromosomes in meiosis I depends on proteolytic cleavage of the meiotic cohesin Rec8 by separin. Cell 103, 387-398. - PubMed
-
- Clyne, R. K., Katis, V. L., Jessop, L., Benjamin, K. R., Herskowitz, I., Lichten, M. and Nasmyth, K. (2003). Polo-like kinase Cdc5 promotes chiasmata formation and cosegregation of sister centromeres at meiosis I. Nat. Cell Biol. 5, 480-485. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

