The crystal structure of the TolB box of colicin A in complex with TolB reveals important differences in the recruitment of the common TolB translocation portal used by group A colicins
- PMID: 19627502
- PMCID: PMC2821528
- DOI: 10.1111/j.1365-2958.2009.06808.x
The crystal structure of the TolB box of colicin A in complex with TolB reveals important differences in the recruitment of the common TolB translocation portal used by group A colicins
Abstract
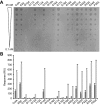
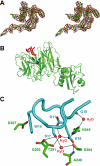
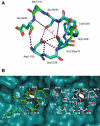
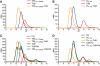
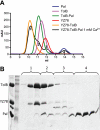
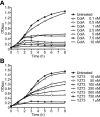
Interaction of the TolB box of Group A colicins with the TolB protein in the periplasm of Escherichia coli cells promotes transport of the cytotoxic domain of the colicin across the cell envelope. The crystal structure of a complex between a 107-residue peptide (TA(1-107)) of the translocation domain of colicin A (ColA) and TolB identified the TolB box as a 12-residue peptide that folded into a distorted hairpin within a central canyon of the beta-propeller domain of TolB. Comparison of this structure with that of the colicin E9 (ColE9) TolB box-TolB complex, together with site-directed mutagenesis of the ColA TolB box residues, revealed important differences in the interaction of the two TolB boxes with an overlapping binding site on TolB. Substitution of the TolB box residues of ColA with those of ColE9 conferred the ability to competitively recruit TolB from Pal but reduced the biological activity of the mutant ColA. This datum explains (i) the difference in binding affinities of ColA and ColE9 with TolB, and (ii) the inability of ColA, unlike ColE9, to competitively recruit TolB from Pal, allowing an understanding of how these two colicins interact in a different way with a common translocation portal in E. coli cells.
Figures

Similar articles
-
Competitive recruitment of the periplasmic translocation portal TolB by a natively disordered domain of colicin E9.Proc Natl Acad Sci U S A. 2006 Aug 15;103(33):12353-8. doi: 10.1073/pnas.0603433103. Epub 2006 Aug 7. Proc Natl Acad Sci U S A. 2006. PMID: 16894158 Free PMC article.
-
Interactions of TolB with the translocation domain of colicin E9 require an extended TolB box.J Bacteriol. 2005 Oct;187(19):6733-41. doi: 10.1128/JB.187.19.6733-6741.2005. J Bacteriol. 2005. PMID: 16166536 Free PMC article.
-
Kinetic basis for the competitive recruitment of TolB by the intrinsically disordered translocation domain of colicin E9.J Mol Biol. 2012 May 18;418(5):269-80. doi: 10.1016/j.jmb.2012.01.039. Epub 2012 Jan 30. J Mol Biol. 2012. PMID: 22310049
-
On mechanisms of colicin import: the outer membrane quandary.Biochem J. 2018 Dec 12;475(23):3903-3915. doi: 10.1042/BCJ20180477. Biochem J. 2018. PMID: 30541793 Review.
-
Colicin A binds to a novel binding site of TolA in the Escherichia coli periplasm.Biochem Soc Trans. 2012 Dec 1;40(6):1469-74. doi: 10.1042/BST20120239. Biochem Soc Trans. 2012. PMID: 23176500 Review.
Cited by
-
Force-Generation by the Trans-Envelope Tol-Pal System.Front Microbiol. 2022 Mar 3;13:852176. doi: 10.3389/fmicb.2022.852176. eCollection 2022. Front Microbiol. 2022. PMID: 35308353 Free PMC article. Review.
-
Interaction of the colicin K bactericidal toxin with components of its import machinery in the periplasm of Escherichia coli.J Bacteriol. 2010 Nov;192(22):5934-42. doi: 10.1128/JB.00936-10. Epub 2010 Sep 24. J Bacteriol. 2010. PMID: 20870776 Free PMC article.
-
Tools and Approaches for Dissecting Protein Bacteriocin Import in Gram-Negative Bacteria.Front Microbiol. 2019 Mar 28;10:646. doi: 10.3389/fmicb.2019.00646. eCollection 2019. Front Microbiol. 2019. PMID: 31001227 Free PMC article. Review.
-
Structural evidence that colicin A protein binds to a novel binding site of TolA protein in Escherichia coli periplasm.J Biol Chem. 2012 Jun 1;287(23):19048-57. doi: 10.1074/jbc.M112.342246. Epub 2012 Apr 9. J Biol Chem. 2012. PMID: 22493500 Free PMC article.
-
The multifarious roles of Tol-Pal in Gram-negative bacteria.FEMS Microbiol Rev. 2020 Jul 1;44(4):490-506. doi: 10.1093/femsre/fuaa018. FEMS Microbiol Rev. 2020. PMID: 32472934 Free PMC article. Review.
References
-
- Abergel C, Bouveret E, Claverie J-M, Brown K, Rigal A, Lazdunski C, Bénédetti H. Structure of the Escherichia coli TolB protein determined by MAD methods at 1.95 Å resolution. Structure. 1999;7:1291–1300. - PubMed
-
- Bainbridge G, Armstrong GA, Dover LG, Whelan KF, Lakey JH. Displacement of OmpF loop 3 is not required for the membrane translocation of colicins N and A in vivo. FEBS Lett. 1998;432:117–122. - PubMed
-
- Bonsor DA, Grishkovskaya I, Dodson EJ, Kleanthous C. Molecular mimicry enables competitive recruitment by a natively disordered protein. J Am Chem Soc. 2007;129:4800–4807. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

