Unnatural imidazopyridopyrimidine:naphthyridine base pairs: selective incorporation and extension reaction by Deep Vent (exo- ) DNA polymerase
- PMID: 19628664
- PMCID: PMC2761277
- DOI: 10.1093/nar/gkp611
Unnatural imidazopyridopyrimidine:naphthyridine base pairs: selective incorporation and extension reaction by Deep Vent (exo- ) DNA polymerase
Abstract
In our previous communication we reported the enzymatic recognition of unnatural imidazopyridopyrimidine:naphthyridine (Im:Na) base pairs, i.e. ImO(N):NaN(O) and ImN(O):NaO(N), using the Klenow fragment exo(-) [KF (exo(-))]. We describe herein the successful results of (i) improved enzymatic recognition for ImN(O):NaO(N) base pairs and (ii) further primer extension reactions after the Im:Na base pairs by Deep Vent DNA polymerase exo(-) [Deep Vent (exo(-))]. Since KF (exo(-)) did not catalyze primer extension reactions after the Im:Na base pair, we carried out a screening of DNA polymerases to promote the primer extension reaction as well as to improve the selectivity of base pair recognition. As a result, a family B DNA polymerase, especially Deep Vent (exo(-)), seemed most promising for this purpose. In the ImO(N):NaN(O) base pair, incorporation of NaN(O)TP against ImO(N) in the template was preferable to that of the natural dNTPs, while incorporation of dATP as well as dGTP competed with that of ImO(N)TP when NaN(O) was placed in the template. Thus, the selectivity of base pair recognition by Deep Vent (exo(-)) was less than that by KF (exo(-)) in the case of the ImO(N):NaN(O) base pair. On the other hand, incorporation of NaO(N)TP against ImN(O) in the template and that of ImN(O)TP against NaO(N) were both quite selective. Thus, the selectivity of base pair recognition was improved by Deep Vent (exo(-)) in the ImN(O):NaO(N) base pair. Moreover, this enzyme catalyzed further primer extension reactions after the ImN(O):NaO(N) base pair to afford a faithful replicate, which was confirmed by MALDI-TOF mass spectrometry as well as the kinetics data for extension fidelity next to the ImN(O):NaO(N) base pair. The results presented in this paper revealed that the ImN(O):NaO(N) base pair might be a third base pair beyond the Watson-Crick base pairs.
Figures





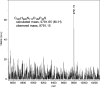

References
-
- Henry AA, Romesberg FE. Beyond A, C, G and T: augmenting nature's alphabet. Curr. Opin. Chem. Biol. 2003;7:727–733. - PubMed
-
- Hirao I. Unnatural base pair systems for DNA/RNA-based biotechnology. Curr. Opin. Chem. Biol. 2006;10:622–627. - PubMed
-
- Hirao I, Mitsui T, Kimoto M, Yokoyama S. An efficient unnatural base pair for PCR amplification. J. Am. Chem. Soc. 2007;129:15549–15555. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

