PARP is activated at stalled forks to mediate Mre11-dependent replication restart and recombination
- PMID: 19629035
- PMCID: PMC2738702
- DOI: 10.1038/emboj.2009.206
PARP is activated at stalled forks to mediate Mre11-dependent replication restart and recombination
Abstract
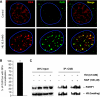
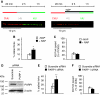
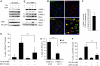
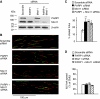
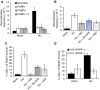
If replication forks are perturbed, a multifaceted response including several DNA repair and cell cycle checkpoint pathways is activated to ensure faithful DNA replication. Here, we show that poly(ADP-ribose) polymerase 1 (PARP1) binds to and is activated by stalled replication forks that contain small gaps. PARP1 collaborates with Mre11 to promote replication fork restart after release from replication blocks, most likely by recruiting Mre11 to the replication fork to promote resection of DNA. Both PARP1 and PARP2 are required for hydroxyurea-induced homologous recombination to promote cell survival after replication blocks. Together, our data suggest that PARP1 and PARP2 detect disrupted replication forks and attract Mre11 for end processing that is required for subsequent recombination repair and restart of replication forks.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


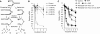


References
-
- Ahel I, Ahel D, Matsusaka T, Clark AJ, Pines J, Boulton SJ, West SC (2008) Poly(ADP-ribose)-binding zinc finger motifs in DNA repair/checkpoint proteins. Nature 451: 81–85 - PubMed
-
- Allinson SL, Dianova II, Dianov GL (2003) Poly(ADP-ribose) polymerase in base excision repair: always engaged, but not essential for DNA damage processing. Acta Biochim Pol 50: 169–179 - PubMed
-
- Ame JC, Rolli V, Schreiber V, Niedergang C, Apiou F, Decker P, Muller S, Hoger T, Menissier-de Murcia J, de Murcia G (1999) PARP-2, a novel mammalian DNA damage-dependent poly(ADP-ribose) polymerase. J Biol Chem 274: 17860–17868 - PubMed
-
- Anachkova B, Russev G, Poirier GG (1989) DNA replication and poly(ADP-ribosyl)ation of chromatin. Cytobios 58: 19–28 - PubMed
-
- Arnaudeau C, Lundin C, Helleday T (2001) DNA double-strand breaks associated with replication forks are predominantly repaired by homologous recombination involving an exchange mechanism in mammalian cells. J Mol Biol 307: 1235–1245 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

