A quantitative systems view of the spindle assembly checkpoint
- PMID: 19629044
- PMCID: PMC2722251
- DOI: 10.1038/emboj.2009.186
A quantitative systems view of the spindle assembly checkpoint
Abstract
The idle assembly checkpoint acts to delay chromosome segregation until all duplicated sister chromatids are captured by the mitotic spindle. This pathway ensures that each daughter cell receives a complete copy of the genome. The high fidelity and robustness of this process have made it a subject of intense study in both the experimental and computational realms. A significant number of checkpoint proteins have been identified but how they orchestrate the communication between local spindle attachment and global cytoplasmic signalling to delay segregation is not yet understood. Here, we propose a systems view of the spindle assembly checkpoint to focus attention on the key regulators of the dynamics of this pathway. These regulators in turn have been the subject of detailed cellular measurements and computational modelling to connect molecular function to the dynamics of spindle assembly checkpoint signalling. A review of these efforts reveals the insights provided by such approaches and underscores the need for further interdisciplinary studies to reveal in full the quantitative underpinnings of this cellular control pathway.
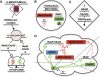
Figures


Similar articles
-
Regulation of APC-Cdc20 by the spindle checkpoint.Curr Opin Cell Biol. 2002 Dec;14(6):706-14. doi: 10.1016/s0955-0674(02)00382-4. Curr Opin Cell Biol. 2002. PMID: 12473343 Review.
-
The spindle checkpoint: tension versus attachment.Trends Cell Biol. 2005 Sep;15(9):486-93. doi: 10.1016/j.tcb.2005.07.005. Trends Cell Biol. 2005. PMID: 16084093 Review.
-
Diverse functions of spindle assembly checkpoint genes in Saccharomyces cerevisiae.Genetics. 2006 Jan;172(1):53-65. doi: 10.1534/genetics.105.046441. Epub 2005 Sep 12. Genetics. 2006. PMID: 16157669 Free PMC article.
-
Protein kinases involved in mitotic spindle checkpoint regulation.Results Probl Cell Differ. 2006;42:93-109. doi: 10.1007/b138827. Results Probl Cell Differ. 2006. PMID: 16903209 Review.
-
Shugoshin ensures maintenance of the spindle assembly checkpoint response and efficient spindle disassembly.Mol Microbiol. 2021 Oct;116(4):1079-1098. doi: 10.1111/mmi.14796. Epub 2021 Aug 30. Mol Microbiol. 2021. PMID: 34407255
Cited by
-
Connecting up and clearing out: how kinetochore attachment silences the spindle assembly checkpoint.Chromosoma. 2012 Oct;121(5):509-25. doi: 10.1007/s00412-012-0378-5. Epub 2012 Jul 11. Chromosoma. 2012. PMID: 22782189 Review.
-
Biophysics of mitosis.Q Rev Biophys. 2012 May;45(2):147-207. doi: 10.1017/S0033583512000017. Epub 2012 Feb 10. Q Rev Biophys. 2012. PMID: 22321376 Free PMC article. Review.
-
Closed MAD2 (C-MAD2) is selectively incorporated into the mitotic checkpoint complex (MCC).Cell Cycle. 2011 Nov 1;10(21):3740-50. doi: 10.4161/cc.10.21.17919. Epub 2011 Nov 1. Cell Cycle. 2011. PMID: 22037211 Free PMC article.
-
p31(comet) acts to ensure timely spindle checkpoint silencing subsequent to kinetochore attachment.Mol Biol Cell. 2011 Nov;22(22):4236-46. doi: 10.1091/mbc.E11-03-0216. Epub 2011 Sep 30. Mol Biol Cell. 2011. PMID: 21965286 Free PMC article.
-
Erroneous Silencing of the Mitotic Checkpoint by Aberrant Spindle Pole-Kinetochore Coordination.Biophys J. 2015 Dec 1;109(11):2418-35. doi: 10.1016/j.bpj.2015.10.024. Biophys J. 2015. PMID: 26636952 Free PMC article.
References
-
- Acquaviva C, Herzog F, Kraft C, Pines J (2004) The anaphase promoting complex/cyclosome is recruited to centromeres by the spindle assembly checkpoint. Nat Cell Biol 22: 22 - PubMed
-
- Buffin E, Lefebvre C, Huang J, Gagou ME, Karess RE (2005) Recruitment of mad2 to the kinetochore requires the rod/zw10 complex. Curr Biol 15: 856–861 - PubMed
-
- Cheeseman IM, Chappie JS, Wilson-Kubalek EM, Desai A (2006) The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell 127: 983–997 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

