Circadian clock genes contribute to the regulation of hair follicle cycling
- PMID: 19629164
- PMCID: PMC2705795
- DOI: 10.1371/journal.pgen.1000573
Circadian clock genes contribute to the regulation of hair follicle cycling
Abstract
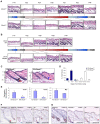
Hair follicles undergo recurrent cycling of controlled growth (anagen), regression (catagen), and relative quiescence (telogen) with a defined periodicity. Taking a genomics approach to study gene expression during synchronized mouse hair follicle cycling, we discovered that, in addition to circadian fluctuation, CLOCK-regulated genes are also modulated in phase with the hair growth cycle. During telogen and early anagen, circadian clock genes are prominently expressed in the secondary hair germ, which contains precursor cells for the growing follicle. Analysis of Clock and Bmal1 mutant mice reveals a delay in anagen progression, and the secondary hair germ cells show decreased levels of phosphorylated Rb and lack mitotic cells, suggesting that circadian clock genes regulate anagen progression via their effect on the cell cycle. Consistent with a block at the G1 phase of the cell cycle, we show a significant upregulation of p21 in Bmal1 mutant skin. While circadian clock mechanisms have been implicated in a variety of diurnal biological processes, our findings indicate that circadian clock genes may be utilized to modulate the progression of non-diurnal cyclic processes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





Similar articles
-
A meeting of two chronobiological systems: circadian proteins Period1 and BMAL1 modulate the human hair cycle clock.J Invest Dermatol. 2014 Mar;134(3):610-619. doi: 10.1038/jid.2013.366. Epub 2013 Sep 4. J Invest Dermatol. 2014. PMID: 24005054
-
Effects of photoperiod on circadian clock genes in skin contribute to the regulation of hair follicle cycling of Inner Mongolia white cashmere goats.Anim Sci J. 2020 Jan-Dec;91(1):e13320. doi: 10.1111/asj.13320. Epub 2019 Dec 16. Anim Sci J. 2020. PMID: 31845459
-
The clock gene brain and muscle Arnt-like protein-1 (BMAL1) is involved in hair growth.Arch Dermatol Res. 2013 Oct;305(8):755-61. doi: 10.1007/s00403-013-1403-0. Epub 2013 Aug 18. Arch Dermatol Res. 2013. PMID: 23955654
-
Clock genes, hair growth and aging.Aging (Albany NY). 2010 Mar 31;2(3):122-8. doi: 10.18632/aging.100130. Aging (Albany NY). 2010. PMID: 20375466 Free PMC article. Review.
-
Plasminogen activator inhibitor-1 and the circadian clock in metabolic disorders.Clin Exp Hypertens. 2009 May;31(3):208-19. doi: 10.1080/10641960902822468. Clin Exp Hypertens. 2009. PMID: 19387897 Review.
Cited by
-
Lef1 expression in fibroblasts maintains developmental potential in adult skin to regenerate wounds.Elife. 2020 Sep 29;9:e60066. doi: 10.7554/eLife.60066. Elife. 2020. PMID: 32990218 Free PMC article.
-
Differential expression of the circadian clock in maternal and embryonic tissues of mice.PLoS One. 2010 Mar 24;5(3):e9855. doi: 10.1371/journal.pone.0009855. PLoS One. 2010. PMID: 20352049 Free PMC article.
-
Overview of the Circadian Clock in the Hair Follicle Cycle.Biomolecules. 2023 Jul 3;13(7):1068. doi: 10.3390/biom13071068. Biomolecules. 2023. PMID: 37509104 Free PMC article. Review.
-
Toward a unified model of developmental timing: A "molting" approach.Worm. 2012 Oct 1;1(4):221-30. doi: 10.4161/worm.20874. Worm. 2012. PMID: 24058853 Free PMC article.
-
Epidermal stem cells ride the circadian wave.Genome Biol. 2013 Nov 29;14(11):140. doi: 10.1186/gb4142. Genome Biol. 2013. PMID: 24286354 Free PMC article.
References
-
- Stenn KS, Paus R. Controls of hair follicle cycling. Physiol Rev. 2001;81:449–494. - PubMed
-
- Paus R, Cotsarelis G. The biology of hair follicles. N Engl J Med. 1999;341:491–497. - PubMed
-
- Muller-Rover S, Handjiski B, van der Veen C, Eichmuller S, Foitzik K, et al. A comprehensive guide for the accurate classification of murine hair follicles in distinct hair cycle stages. J Invest Dermatol. 2001;117:3–15. - PubMed
-
- Millar SE. Molecular mechanisms regulating hair follicle development. J Invest Dermatol. 2002;118:216–225. - PubMed
-
- Fuchs E, Merrill BJ, Jamora C, DasGupta R. At the roots of a never-ending cycle. Dev Cell. 2001;1:13–25. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

