Significant bias against the ACA triplet in the tmRNA sequence of Escherichia coli K-12
- PMID: 19633073
- PMCID: PMC2747887
- DOI: 10.1128/JB.00699-09
Significant bias against the ACA triplet in the tmRNA sequence of Escherichia coli K-12
Abstract
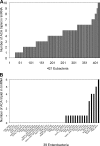
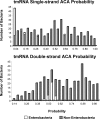
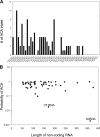
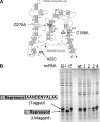
The toxin MazF in Escherichia coli cleaves single-stranded RNAs specifically at ACA sequences. MazF overexpression virtually eliminates all cellular mRNAs to completely block protein synthesis. However, protein synthesis can continue on an mRNA that is devoid of ACA triplets. The finding that ribosomal RNAs remain intact in the face of complete translation arrest suggested a purpose for such preservation. We therefore examined the sequences of all transcribed RNAs to determine if there was any statistically significant bias against ACA. While ACA motifs are absent from tmRNA, 4.5S RNA, and seven of the eight 5S rRNAs, statistical analysis revealed that only for tmRNA was the absence nonrandom. The introduction of single-strand ACAs makes tmRNA highly susceptible to MazF cleavage. Furthermore, analysis of tmRNA sequences from 442 bacteria showed that the discrimination against ACA in tmRNAs was seen mostly in enterobacteria. We propose that the unusual bias against ACA in tmRNA may have coevolved with the acquisition of MazF.
Figures



References
-
- Bjork, G. R. 1996. Stable RNA modification, vol. 2. ASM Press, Washington, DC.
-
- Christensen, S. K., and K. Gerdes. 2003. RelE toxins from bacteria and archaea cleave mRNAs on translating ribosomes, which are rescued by tmRNA. Mol. Microbiol. 48:1389-1400. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

