Genome-wide identification of Saccharomyces cerevisiae genes required for maximal tolerance to ethanol
- PMID: 19633105
- PMCID: PMC2747848
- DOI: 10.1128/AEM.00845-09
Genome-wide identification of Saccharomyces cerevisiae genes required for maximal tolerance to ethanol
Abstract
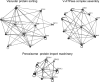
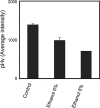
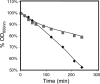
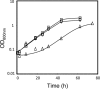
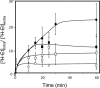
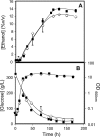
The understanding of the molecular basis of yeast resistance to ethanol may guide the design of rational strategies to increase process performance in industrial alcoholic fermentations. In this study, the yeast disruptome was screened for mutants with differential susceptibility to stress induced by high ethanol concentrations in minimal growth medium. Over 250 determinants of resistance to ethanol were identified. The most significant gene ontology terms enriched in this data set are those associated with intracellular organization, biogenesis, and transport, in particular, regarding the vacuole, the peroxisome, the endosome, and the cytoskeleton, and those associated with the transcriptional machinery. Clustering the proteins encoded by the identified determinants of ethanol resistance by their known physical and genetic interactions highlighted the importance of the vacuolar protein sorting machinery, the vacuolar H(+)-ATPase complex, and the peroxisome protein import machinery. Evidence showing that vacuolar acidification and increased resistance to the cell wall lytic enzyme beta-glucanase occur in response to ethanol-induced stress was obtained. Based on the genome-wide results, the particular role of the FPS1 gene, encoding a plasma membrane aquaglyceroporin which mediates controlled glycerol efflux, in ethanol stress resistance was further investigated. FPS1 expression contributes to decreased [(3)H]ethanol accumulation in yeast cells, suggesting that Fps1p may also play a role in maintaining the intracellular ethanol level during active fermentation. The increased expression of FPS1 confirmed the important role of this gene in alcoholic fermentation, leading to increased final ethanol concentration under conditions that lead to high ethanol production.
Figures

References
-
- Aguilera, F., R. A. Peinado, C. Millan, J. M. Ortega, and J. C. Mauricio. 2006. Relationship between ethanol tolerance, H+-ATPase activity and the lipid composition of the plasma membrane in different wine yeast strains. Int. J. Food Microbiol. 110:34-42. - PubMed
-
- Alexandre, H., V. Ansanay-Galeote, S. Dequin, and B. Blondin. 2001. Global gene expression during short-term ethanol stress in Saccharomyces cerevisiae. FEBS Lett. 498:98-103. - PubMed
-
- Alper, H., J. Moxley, E. Nevoigt, G. R. Fink, and G. Stephanopoulos. 2006. Engineering yeast transcription machinery for improved ethanol tolerance and production. Science 314:1565-1568. - PubMed
-
- Carmelo, V., H. Santos, and I. Sá-Correia. 1997. Effect of extracellular acidification on the activity of plasma membrane ATPase and on the cytosolic and vacuolar pH of Saccharomyces cerevisiae. Biochim. Biophys. Acta 1325:63-70. - PubMed
-
- Casey, G. P., and W. M. Ingledew. 1986. Ethanol tolerance in yeasts. Crit. Rev. Microbiol. 13:219-280. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

